Introduction
Blood is approximately 99% red blood cells (RBCs) and only ~1% white blood cells (WBCs), yet RNA of interest resides almost exclusively in WBCs. That means you're working with a small target fraction buried in a large, chemically hostile background.
RBCs contain high concentrations of RNases that release during lysis or freeze-thaw cycles and rapidly degrade RNA if not immediately inactivated. Hemoglobin compounds the problem further, acting as a potent inhibitor of RT-PCR and reverse transcriptase enzymes while physically clogging silica spin columns and magnetic bead matrices.
Choosing the wrong collection tube, storage method, or kit has tangible consequences: degraded RNA with low RIN scores that are unsuitable for RNA-seq, RT-PCR, or transcriptomic biomarker studies. The cost is real — wasted samples, repeated extractions, and unreliable data.
For mRNA-seq library construction, most providers including Illumina require RIN ≥ 8. Low-RIN RNA skews gene expression profiles and increases false-negative rates in sequencing.
What follows is a protocol-by-protocol comparison of blood RNA extraction methods, stabilisation strategies, and a practical kit selection framework — covering everything from fresh whole blood to frozen legacy samples.
TL;DR
- Fresh blood processed immediately gives the best RNA yield and quality; avoid freezing in plain EDTA tubes
- RNA stabilisation tubes (PAXgene, Tempus) preserve integrity at room temperature for up to 5 days
- For frozen EDTA legacy samples, adding lysis buffer during thawing (not after) can rescue RNA to RIN > 7
- Match kit selection to sample logistics, target RIN, throughput, and downstream application
- High-throughput labs benefit from automated extraction platforms to reduce variability and boost reproducibility
Why RNA Isolation from Blood is Uniquely Challenging
The Composition Problem
Whole blood is approximately 4.3–6.2 million RBCs per µL, 150,000–400,000 platelets per µL, and only 4,500–11,000 WBCs per µL. Leukocytes (WBCs) comprise less than 1% of the cellular blood fraction but contain the primary RNA of interest for transcriptomics. Extraction workflows must therefore separate a tiny target population from an overwhelming background of RBCs and platelets — which is why blood-specific protocols exist rather than generic RNA extraction approaches.
RNase Contamination as a Primary Failure Mode
Human serum contains high levels of Ribonuclease 1 (RNase 1), circulating at approximately 250 ng/mL. RBCs harbour even higher concentrations of RNases that are released during RBC lysis steps or freeze-thaw cycles. Without immediate inactivation using guanidine-containing buffers or proprietary stabilisation reagents, these RNases rapidly degrade RNA within minutes.
Hemoglobin Interference
Hemoglobin from RBCs is a potent inhibitor of molecular reactions. Hematin (a heme derivative) causes complete amplification inhibition in qPCR at 80 µM and in digital PCR at 3000 µM. It also physically clogs silica spin column filters and magnetic bead matrices, reducing binding capacity, yield, and purity.
Poor A260/230 ratios in blood RNA extractions are frequently traceable to this source — residual hemoglobin contaminating the eluate rather than salt carryover or phenol interference.
The Freeze-Thaw Degradation Cascade
In conventional EDTA tubes (which contain no RNA stabilisers), freezing temporarily deactivates RNases. However, thawing reactivates them in an uncontrolled environment alongside ruptured WBCs, causing rapid RNA degradation. RIN typically falls below 4–5 using traditional methods on frozen EDTA blood.
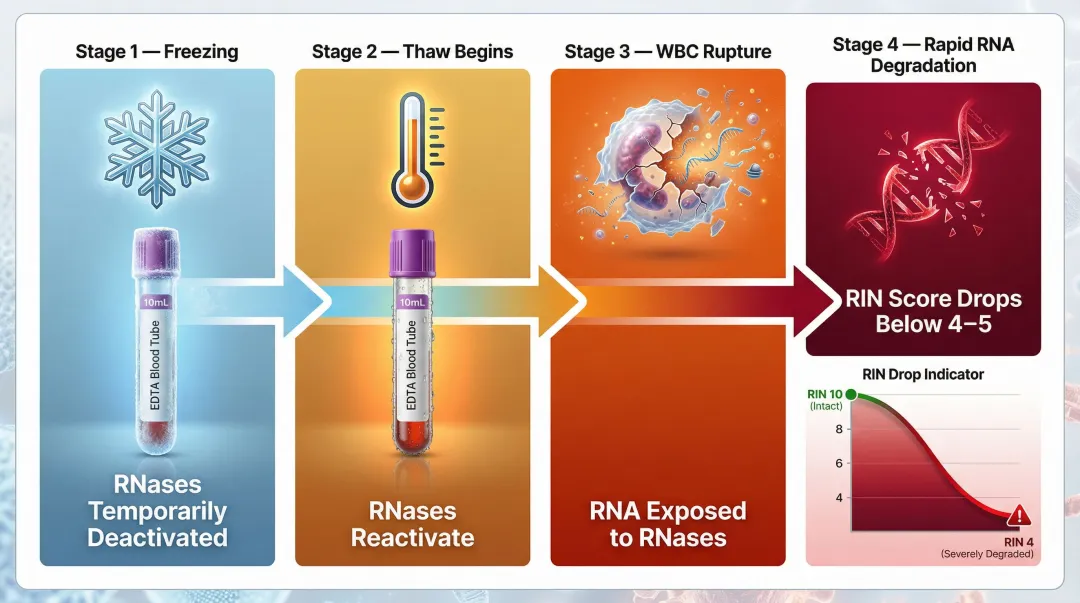
The RIN Benchmark That Matters
That RIN drop matters more than most researchers expect. Illumina recommends a minimum RIN of 8 for its TruSeq Stranded mRNA kit (poly-A selection). RNA below this threshold risks incomplete library construction, skewed expression profiles, and false-negative results. Low RNA integrity (RIN < 6) causes a 30% loss of short to mid-sized transcripts and introduces severe 3' coverage bias in poly-A selected libraries.
The five challenges above — cellular composition, RNase activity, hemoglobin inhibition, freeze-thaw instability, and RIN requirements — each demand specific protocol decisions. The sections below map those decisions to the collection methods and extraction kits best suited to address them.
Blood Collection, Storage, and Pre-Extraction Best Practices
Fresh Blood: Optimal Starting Material
Fresh blood processed within ~1 hour of collection produces the highest RNA yield and integrity. If immediate processing isn't possible, brief storage at 4°C (not frozen) for up to a few hours is acceptable. Never freeze in a plain EDTA tube if intact RNA is the goal.
Intermediate "Stop Point" Strategy:Perform RBC lysis on fresh blood, pellet WBCs, then lyse the WBC pellet in a guanidine-containing buffer (which inactivates RNases) and store at −20°C. This allows processing to pause safely without RNA degradation.
Stabilisation Tube Systems
When fresh processing isn't feasible, stabilisation tube systems extend the pre-extraction window without compromising RNA integrity.
PAXgene Blood RNA Tube:Blood is collected into a tube pre-filled with proprietary RNA stabilisation buffer that immediately lyses cells and precipitates RNA, protecting it from RNase activity. Samples can be stored at room temperature for up to 5 days before purification, making it well-suited for multi-site clinical studies.
Note: a minimum 2-hour room temperature incubation is required after collection before processing or freezing.
Tempus Blood RNA Tube:A similar vacutainer-based stabilisation approach from ThermoFisher that collects 3 mL of blood. Samples can be stored at room temperature for up to 5 days or at 4°C for at least 7 days. The key advantage: no minimum incubation time before processing (unlike PAXgene's 2-hour requirement). However, specialised equipment may be required for the purification step.
Frozen EDTA Blood: The Rescue Approach
Research published in Scientific Reports (2024) shows that RNA degradation from frozen EDTA blood occurs primarily during thawing, not during cryopreservation itself. Adding RNA stabilisation or cell lysis buffer directly to the frozen EDTA tube while it is still frozen (before any thawing begins) captures RNases before they reactivate. This approach improves RIN from below 5 to above 7 and increases yield up to 5-fold compared to post-thaw addition.
Protocol and Kit Comparison for Blood RNA Extraction
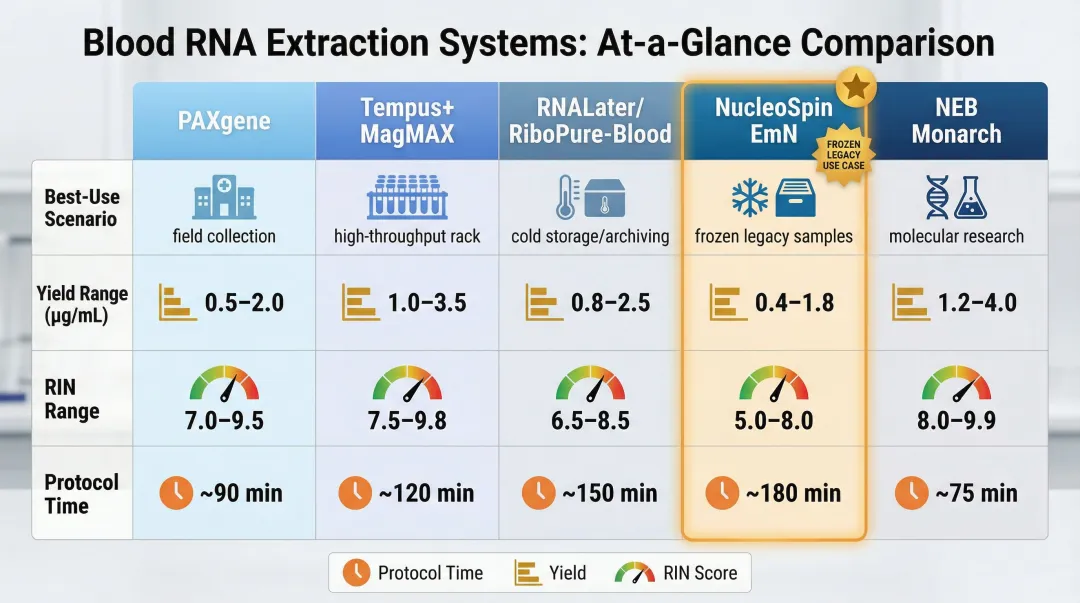
The right choice depends on whether blood is fresh, freshly stabilised, or historically frozen in EDTA tubes. The table below summarises the five major systems, followed by detailed protocol and performance breakdowns for each.
| System | Best For | Yield (µg/mL) | RIN Range | Protocol Time |
|---|---|---|---|---|
| PAXgene Blood RNA System | Fresh blood, standardised biobanking | ~3.1 | 6.0–8.6 | ~2 hours |
| Tempus + MagMAX | Fresh blood, higher yield priority | 2.0–8.0 | 7.1–8.9 | Instrument-dependent |
| RNALater/RiboPure-Blood | Small-volume fresh samples | 4.0–8.0 | 4.6–5.8 | ~45 minutes |
| Nucleospin EmN Protocol | Frozen EDTA legacy samples | 7.0–9.0 | ~7–8 | ~45 minutes |
| NEB Monarch Miniprep | Fresh or frozen, scalable input | 2.5–5.0 | 7.0–8.0 | ~60 minutes |

PAXgene Blood RNA System (BD/PreAnalytix)
Workflow:
- Blood collected in PAXgene tubes is centrifuged to pellet precipitated nucleic acids, washed, and purified using a spin column kit
- Full protocol involves ~21 steps and can take up to 2 hours
- Minimum 2-hour room temperature incubation required before processing
- DNA contamination has been reported in some studies
Performance:
Tempus Blood RNA Tube + MagMAX Kit
Workflow:
- No minimum incubation time required (advantage over PAXgene)
- Paired with MagMAX extraction kit (magnetic bead-based)
- Dedicated instrument (ABI PRISM 6100 or equivalent) required for recommended purification workflow, adding capital cost
Performance:
RNALater/RiboPure-Blood (Ambion/ThermoFisher)
Workflow:
- RNALater added directly to 0.3–0.5 mL of blood to stabilise RNA for up to 3 days at room temperature
- Dual-purification: initial phenol/chloroform extraction (removes hemoglobin and proteins) followed by glass fibre spin column
- Shortest protocol (~45 min total including DNase step) but small input volume limits total yield
Performance:
Nucleospin Blood RNA Kit (Macherey-Nagel) with EDTA Mixed-Thawing Protocol (EmN)
Workflow:
- Particularly valuable for frozen EDTA legacy samples
- Nucleospin lysis buffer added directly to frozen EDTA tube; sample thaws in presence of RNase inhibitors
- After mixing, ~1.3 mL proceeds through Nucleospin RNA workflow (8 steps, ~45 minutes)
Performance:
- Achieves RIN comparable to PAXgene control (~7–8) but with up to 5× higher RNA yield
- Typical yield: 7.0–9.0 µg per mL blood
- Purity (A260/280): 1.9–2.1
- Enables simultaneous DNA extraction from remaining lysate—directly useful for studies requiring matched DNA/RNA from a single sample
NEB Monarch Total RNA Miniprep Kit
Workflow:
- Flexible for both fresh and frozen blood
- Processes up to 200 µL with standard protocol or up to 3 mL with column reloading
- Requires Monarch DNA/RNA Protection Reagent as 2X concentrate added during thawing of frozen samples
- Skips gDNA removal column in whole blood mode; on-column DNase I treatment recommended
Performance:
- Typical yield: 2.5–5.0 µg per mL blood
- Achievable RIN: 7.0–8.0
- Purity (A260/280): 2.0
- Well-suited for labs that process variable input volumes without committing to a dedicated collection tube system
Each system above addresses a distinct workflow scenario. The next step is matching these profiles to your sample type, throughput requirements, and downstream application — covered in the selection guide below.
Key Factors to Consider When Selecting a Blood RNA Extraction Protocol
No single kit is optimal for all use cases. These factors help labs match technical specifications to their actual workflow needs.
Sample Logistics and Collection Site
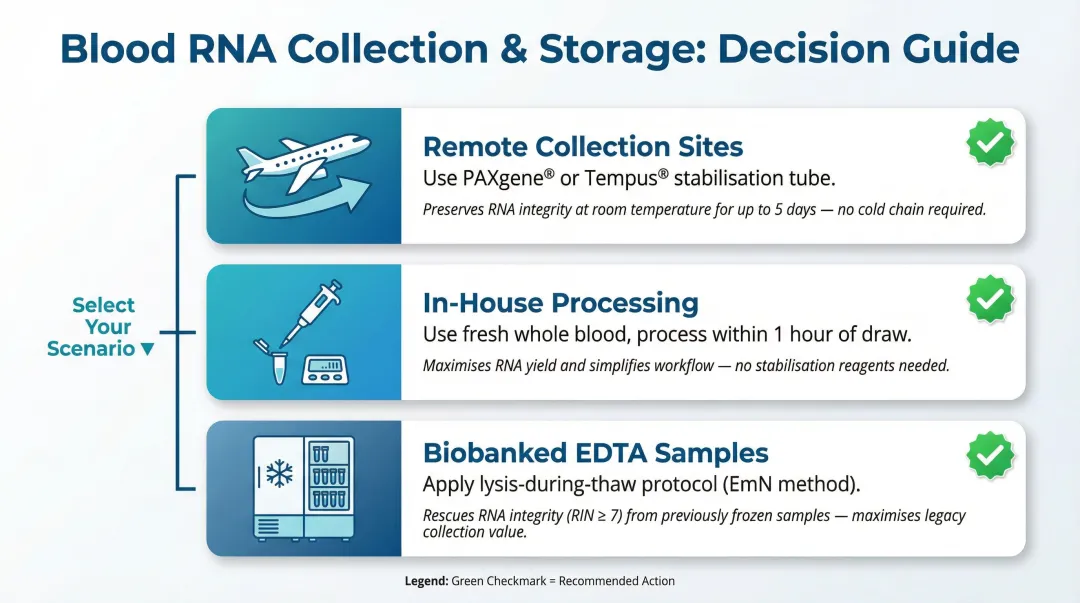
Collection logistics often determine protocol choice before any technical comparison begins:
- Remote collection sites — Stabilisation tube systems (PAXgene or Tempus) are the only reliable option when blood is transported before processing. RNA integrity degrades rapidly without chemical stabilisation.
- In-house processing — Fresh blood protocols are preferred when samples are processed within ~1 hour of draw. They deliver higher yield with simpler workflows.
- Biobanked EDTA samples — The EmN (EDTA-mixed Nucleospin) or similar lysis-during-thaw approach is the only viable path to high-quality RNA from frozen EDTA samples.

Downstream Application Requirements
The minimum acceptable RNA quality varies by application. Match your protocol accordingly:
| Application | Minimum RIN | Notes |
|---|---|---|
| RNA-seq (mRNA-seq) | ≥ 8 | Illumina TruSeq Stranded mRNA strictly requires RIN ≥ 8 |
| RT-qPCR | ≥ 6 | Higher RIN improves reproducibility |
| Microarray analysis | ≥ 6 | Accuracy improves with higher integrity |
Yield matters too. If 500 ng is the minimum input for library prep, kits with lower recovery — such as PAXgene-based systems used on small blood volumes — may require larger input volumes or multiple extractions.
Cost, Throughput, and Automation Readiness
Lab scale and workflow structure determine which cost and throughput tradeoffs are acceptable.
For low-throughput research labs, cost-per-prep and hands-on time dominate the decision. Manual column-based kits (Nucleospin, NEB Monarch) offer flexibility without capital equipment costs.
For high-throughput clinical or biobanking settings, batch capacity and system integration define throughput limits. Automated extraction platforms reduce hands-on time from 75 minutes to 20 minutes for 24 samples, while also improving inter-operator reproducibility.
On automation compatibility:
- Phenol/chloroform kits (such as RiboPure-Blood) are poorly suited to automated liquid handling
- Magnetic bead-based protocols (MagMAX, Nucleospin) integrate directly with most liquid handling platforms
- Column-based kits sit in between — automatable with the right deck setup, but less seamless than bead-based systems
When calculating total cost, factor in tube cost, kit cost, instrument requirements, and labour time — not just per-prep reagent price.
How Cambrian Bioworks Can Help Streamline Blood RNA Extraction
Cambrian Bioworks provides compact, automated nucleic acid extraction solutions built for molecular diagnostics and research labs that need consistent, scalable results. Their systems are designed to remove variability from manual extraction workflows—a critical need when working with challenging samples like blood-derived RNA.
The Beluga automated liquid handler and **Manta CE-IVD certified extraction system** are open-system compatible, configurable to run established bead-based RNA extraction chemistries. This lets labs standardise protocols and reduce hands-on time without being locked into a single vendor's reagents.
For labs wanting a fully integrated option, the Cambrian Blood RNA Isolation Kit—optimised for the Manta platform—delivers RIN values of 8.6–9.7 from whole blood, with scalable yields of 2–10 µg per sample depending on input volume. Key workflow advantages include:
- Extraction completed in ~30 minutes with no batching pressure
- Open-system compatibility with established bead-based chemistries
- Minimal genomic DNA contamination in RNA eluates
- Scalable throughput without protocol revalidation

Backed by ISO 13485 certification and a CE-IVD mark, Cambrian's platforms are trusted by 67+ labs globally—including Tata Memorial Hospital, ACTREC, and Strand Life Sciences. Labs scaling RNA extraction for biomarker studies, oncology diagnostics, or infectious disease research can automate without replacing existing validated protocols, keeping workflows intact while increasing throughput.
Frequently Asked Questions
What does EDTA do to RNA?
EDTA itself does not stabilise RNA. It chelates calcium and magnesium ions to prevent blood clotting, but does not inhibit RNases, which are metal-ion-independent enzymes. Without an RNA stabilisation buffer, EDTA-collected blood—especially after freezing—releases active RNases upon lysis or thawing, leading to significant RNA degradation.
How to store whole blood for RNA extraction?
Three main options:
- Process fresh blood within 1 hour; store at 4°C only for brief periods
- Use a stabilisation tube (PAXgene or Tempus) for room temperature storage up to 5 days
- If freezing is unavoidable, add RNA stabilisation or lysis buffer during thawing to limit RNase reactivation
What RIN score is required for RNA-seq from blood?
Most mRNA-seq library preparation protocols and sequencing providers (including Illumina) recommend a minimum RIN of 7–8. RNA below this threshold risks incomplete library construction, skewed expression profiles, and false-negative sequencing results. Total RNA-seq can tolerate slightly lower RIN but at the cost of accuracy for coding RNA detection.
Can you extract RNA and DNA simultaneously from the same blood sample?
Yes. Some kits allow co-extraction from the same lysate — one aliquot proceeds to RNA purification while the remainder goes to DNA extraction. This is especially useful for frozen EDTA samples where available blood volume is limited.
How does hemoglobin interfere with RNA isolation from blood?
Hemoglobin is a potent inhibitor of RT-PCR and reverse transcriptase, and physically clogs silica spin column membranes and bead matrices, reducing binding capacity and yield. Protocols using a phenol/chloroform extraction step (such as RiboPure-Blood) remove hemoglobin prior to column purification, improving both purity and downstream performance.
What is the difference between PAXgene and Tempus blood collection tubes for RNA?
Both systems lyse blood cells and inactivate RNases on collection, enabling room temperature storage for up to 5 days. Key differences: PAXgene collects 2.5 mL and requires a 2-hour incubation before processing; Tempus collects 3 mL with no minimum incubation. Instrument requirements and cost per tube also vary, affecting suitability based on lab infrastructure and study design.