Introduction
When a lab processing hundreds of blood samples per run gets degraded RNA, the downstream consequences aren't minor — RT-qPCR fails, transcriptomics data becomes unreliable, and clinical results can't be reported. High-throughput RNA extraction from blood (the large-scale, automated isolation of RNA from whole blood, PBMCs, or leukocyte fractions) is where those failures start or get prevented.
The global molecular diagnostics market was valued at $18.85 billion in 2025 and is projected to reach $25.59 billion by 2033, reflecting the growing pressure on labs to scale molecular workflows without sacrificing quality.
This guide covers what makes blood a uniquely difficult matrix for RNA extraction, how automation addresses those challenges, and what quality benchmarks labs should hold workflows to at scale.
TLDR
- High-throughput RNA extraction from blood automates large-scale RNA isolation for molecular diagnostics, transcriptomics, and liquid biopsy workflows
- Blood presents unique challenges including heme-based PCR inhibition, high endogenous RNase activity, and anticoagulant interference
- Magnetic bead-based workflows are preferred over filter-plate methods for superior scalability, consistency, and automation compatibility
- RNA quality is assessed using RIN scores, A260/A280, and A260/A230 ratios, with stricter inhibitor controls required for blood-derived RNA
- Automation may not suit labs with low, irregular sample volumes or highly variable input matrices
What Is High-Throughput RNA Extraction from Blood?
High-throughput RNA extraction from blood is a scaled, automated workflow that isolates total or messenger RNA from blood samples in parallel. Most platforms process 24 to 96+ samples per run while maintaining RNA yield and integrity sufficient for downstream molecular analysis.
The goal is consistent, inhibitor-free RNA at concentrations suitable for:
- RT-qPCR and quantitative gene expression profiling
- RNA sequencing (RNA-seq) and transcriptome analysis
- Liquid biopsy applications, including circulating RNA detection
Unlike standard single-tube extraction, high-throughput formats use 96-well plates, automated liquid handlers, and pre-validated reagent chemistries adapted specifically for blood matrices. Manual column-based methods handle one to eight samples at a time with considerable operator variability. Automation standardizes lysis buffer addition, incubation timing, and washing steps across entire batches — eliminating the inconsistencies that compound across large sample sets.
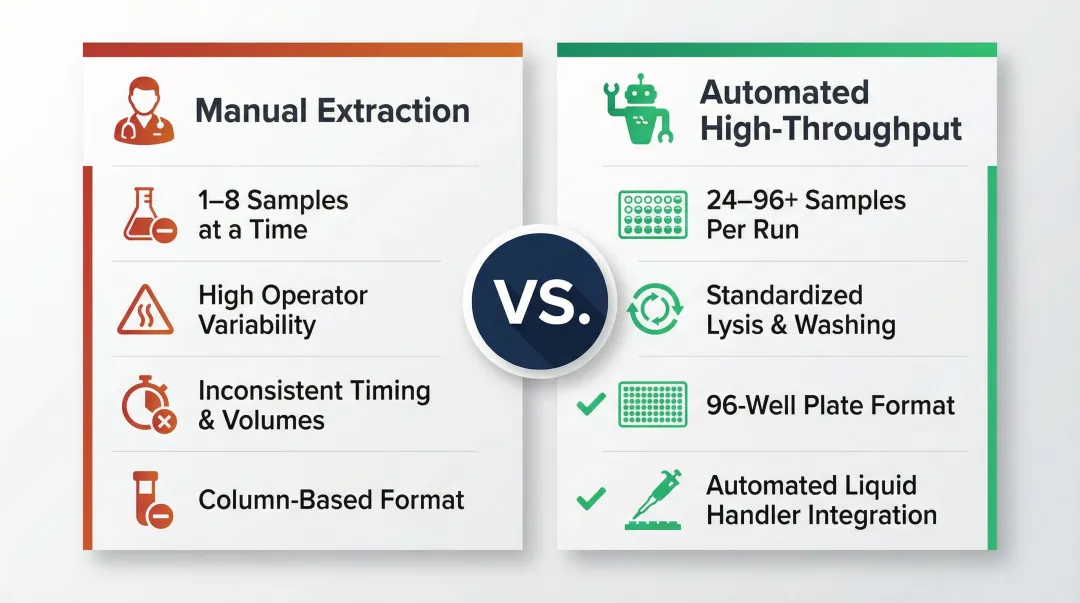

Why Automated High-Throughput RNA Extraction Matters in Blood-Based Labs
Scale Demands Drive Automation
Clinical and research labs running transcriptomic panels process over 25,000 whole transcriptomes annually using advanced automated platforms. The growth in RNA sequencing workflows has driven demand for reproducible, high-volume extraction. High-throughput sequencing platforms like the Illumina NovaSeq 6000 output up to 6 Tb and 20 billion single reads in under two days, which makes sample preparation the rate-limiting step.
Blood-Specific Challenges Require Specialized Solutions
Blood-based workflows introduce challenges not seen in other sample types: active RNase suppression from the point of collection, hemoglobin and heme as potent PCR inhibitors, compatibility with multiple anticoagulant tube types, and reproducibility across variable white blood cell counts.
Blood plasma contains extremely high ribonuclease activity, making immediate stabilization critical. Hemoglobin and immunoglobulin G inhibit DNA polymerase activity, while hematin causes complete amplification inhibition at 3,000 µM in digital PCR.
These chemistry-level variables compound under manual processing. At scale, labs lose control of each factor individually — and the downstream effects are measurable:
- Operator-to-operator variability in RNA yield and purity
- Inconsistent lysis and washing steps leading to inhibitor carryover
- RNA degradation from extended processing times
- Batch failures requiring sample re-collection (often impossible in clinical settings)
Regulatory Requirements Make Automation Essential
For clinical IVD-grade workflows, reproducibility and traceability are regulatory requirements, not aspirational targets. Key obligations include:
- ISO 13485:2016 Clause 7.5.6: Mandates validation of processes where output cannot be verified by subsequent monitoring
- ISO 13485:2016 Clause 7.5.9: Requires documented procedures for traceability across the extraction workflow
- EU IVDR 2017/746, Annex I Section 9: Requires devices to achieve specified analytical performance — sensitivity, specificity, precision, and limits of detection
Automated extraction systems help labs meet these standards consistently through built-in run logging, barcode tracking, and process validation.
How High-Throughput RNA Extraction from Blood Works
The workflow is a multi-stage process beginning at blood collection and ending with quantified, quality-assessed RNA ready for downstream use. Each stage is a control point where failures compound downstream.
Two main chemistries dominate: silica-based filter plate methods (vacuum or centrifuge-driven) and magnetic bead-based methods. Magnetic bead methods are increasingly preferred in automated workflows due to easier liquid handler integration, lower clog risk, and more consistent recovery from blood lysates.
Sample Collection and Pre-Processing
Blood collection tube choice critically determines RNA quality before any extraction step. PAXgene Blood RNA Tubes stabilize intracellular RNA for up to 3 days at 18–25°C, up to 5 days at 2–8°C, and up to 11 years at -20°C or -70°C. Tempus Blood RNA Tubes stabilize RNA for up to 5 days at room temperature and at least 7 days at 4°C.
In contrast, RNA in standard EDTA tubes degrades rapidly—significant integrity decline occurs after 6 hours at room temperature, with RIN values dropping below 5 after 24-48 hours. Heparin tubes should be avoided; heparin inhibits both MMLV reverse transcriptase and Taq DNA polymerase, though this inhibition is reversible by treating RNA with 1-3 units of heparinase I prior to reverse transcription.
Pre-processing differs by tube type: EDTA tubes typically require lysis of red blood cells to isolate leukocytes (the RNA-rich fraction) or direct whole blood lysis using chaotropic buffer. PAXgene/Tempus tubes undergo different lysis protocols post-centrifugation. Automated liquid handlers standardize lysis buffer addition, incubation timing, and mixing — removing a key source of batch-to-batch variability.
Binding and RNA Capture
After lysis, RNA is selectively captured onto silica membrane filters (in high-salt, low-pH buffers) or magnetic beads (which bind RNA through electrostatic or affinity interactions). In bead-based formats, magnetic beads disperse in the lysate, incubate with mixing for full contact, then are pulled to the plate wall by a magnetic plate. The supernatant containing inhibitors, proteins, heme, and lipids is aspirated away. Bead-to-sample ratio and bead resuspension quality are critical control variables at this stage.
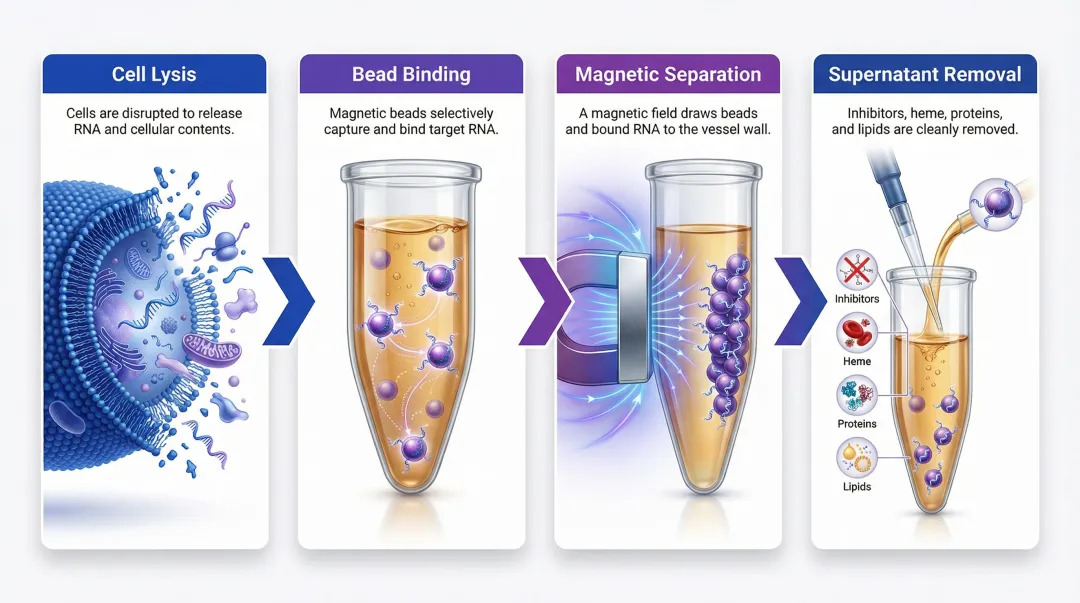
Controlling these variables consistently at scale is where automation earns its place. Cambrian Bioworks' Beluga automated liquid handler integrates validated magnetic bead extraction chemistries with flexible throughput from 1 to 32 samples per run — in a compact, hood-compatible footprint suited to clinical settings.
Washing, Elution, and Quality Assessment
Two to three sequential washes with ethanol-containing buffers remove residual heme pigments, protein fragments, and salt carryover. Automation ensures consistent wash volumes and timing, reducing inhibitor carryover—the primary cause of downstream RT-qPCR failures in blood RNA extracts. Elution is performed with low-salt buffer or nuclease-free water, with eluate volume optimized for downstream assay input concentration.
Quality assessment includes:
- RNA yield: ng/µL via spectrophotometry or fluorometry
- Purity ratios: A260/A280 of ~2.0 indicates pure RNA; A260/A230 >1.8 indicates low salt/solvent carryover
- RNA integrity: RIN scores via Bioanalyzer or Fragment Analyzer
For RNA-Seq, Illumina recommends RIN ≥ 8 for optimal results, though the GTEx project accepts RIN ≥ 6.0. DV200 (percentage of RNA fragments >200 nucleotides) is a more reliable predictor of successful library prep for degraded samples than RIN; Illumina recommends DV200 > 70% for high quality.
Blood RNA typically has lower RIN values than cell-line RNA, largely due to globin mRNA dominance. Reticulocytes contain large amounts of hemoglobin mRNA (HBA1, HBA2, HBB), representing 52–76% of total RNA and consuming disproportionate sequencing reads.
This makes globin depletion a practical necessity for RNA-Seq workflows. Without depletion, hemoglobin genes account for 10–33% of raw gene counts; targeted depletion methods such as Globin-Zero can reduce this to less than 0.1%.
Key Factors That Affect RNA Quality in High-Throughput Blood Workflows
Pre-Analytical Variables and Input Material
Blood volume, collection-to-processing time, storage temperature, and freeze-thaw history are among the most impactful variables on RNA yield and integrity. The evidence for tight handling controls is clear:
- A 2-hour delay at room temperature in an EDTA tube significantly degrades RNA
- PAXgene Blood RNA Tubes tolerate up to two freeze/thaw cycles without significant impact
- Freeze-thawing un-stabilized EDTA blood causes hemolysis that severely degrades RNA, with 28S/18S ratios dropping below 1.0
Inhibitor Load from the Blood Matrix
The blood matrix introduces potent PCR inhibitors that directly affect downstream assay performance. Heme and hemoglobin strongly inhibit both reverse transcriptase and DNA polymerase, and heme inhibits HIV-1 reverse transcriptase in a noncompetitive manner. A visible pink or red tint in the eluate is a reliable predictor of assay failure.

Washing stringency and bead wash volume must be optimized specifically for blood matrices to clear residual heme effectively.
Extraction Chemistry and Automation Compatibility
Not all RNA extraction chemistries are validated for automated pipetting. Viscosity, reagent foaming, and tip-wetting behavior differ meaningfully between manual and automated liquid handlers — and a chemistry that performs well by hand may produce inconsistent yields on a robotic platform.
Labs should use chemistries with verified automation protocols for their specific instrument. For example, Cambrian Bioworks' Beluga automated liquid handler is designed to fit inside a biosafety hood and integrates with validated extraction chemistries suited to clinical environments.
Scale and Batch Size Considerations
High-throughput setups require tight consistency across 96-well formats. Temperature uniformity during lysis and incubation affects bead binding kinetics, and even small variation across plate positions can skew results at scale.
Labs should validate their instrument's pipetting accuracy at blood lysis volumes (typically 100–300 µL) and confirm CV% across replicates before moving to production runs.
Regulatory and Quality Requirements
For IVD-classified workflows, extraction platforms must meet defined performance thresholds across sensitivity, specificity, and repeatability. ISO 13485 and EU IVDR frameworks require documented performance evaluation reports, and India's CDSCO mandates Performance Evaluation Reports for Class B, C, and D IVDs.
Automated systems with barcoded tube compatibility and run logging directly support these traceability requirements.
Common Pitfalls, Misconceptions, and When Automation May Not Be the Right Fit
A common assumption in blood RNA workflows is that high RIN scores confirm extraction quality. They don't. Heme carryover can suppress RT-qPCR even when RNA integrity looks acceptable on a Bioanalyzer. The MIQE guidelines are explicit: spectrophotometric A260/A280 ratios do not detect PCR inhibitors; inhibition must be assessed via sample dilution or universal inhibition assays. The External RNA Controls Consortium provides synthetic RNA spike-in controls that can be introduced to monitor amplification, detect PCR inhibitors, and assess RNA-Seq library prep performance.
A related mistake is adopting 96-well extraction formats optimized for cell culture or swab samples without re-validating for blood. Lysis buffer volumes, incubation times, and wash stringency all behave differently with blood's higher lipid and protein load. Skipping re-validation before clinical deployment risks systematic extraction failures that standard QC metrics won't catch.
When Automation May Not Be the Right Fit
Automation is not the right answer for every lab. Three scenarios where manual or semi-automated approaches may be more practical:
- Low daily volume: Labs processing fewer than 20–30 samples per day may not achieve sufficient ROI. Automated RNA extraction can cost up to 3.5× more for consumables than manual extraction; manual methods are often preferred in resource-limited or low-throughput settings.
- Unpredictable sample load: Labs with highly irregular intake may find manual or semi-automated extraction offers more scheduling flexibility.
- Extremely low input volume: For paediatric or rare disease applications with <100 µL blood, standard RNA-Seq protocols frequently fail due to low WBC counts (< 0.1 × 10⁹/L), with up to 32% of samples yielding insufficient RNA. Specialized ultra-low input kits enabling full-length cDNA synthesis from as little as 2 pg to 10 ng of total RNA are required in these cases.

For labs handling FFPE or tissue-based RNA alongside blood, unified extraction platforms require separate validation for each matrix type — performance in one does not transfer to the other. Extraction-free direct RT-qPCR has been explored for high-volume screening, but lacks the breadth of validation needed for routine clinical blood RNA diagnostics across diverse gene targets.
Frequently Asked Questions
What blood collection tubes are best for high-throughput RNA extraction?
PAXgene and Tempus tubes are preferred for gene expression work because they stabilize RNA immediately at collection. EDTA tubes are acceptable for same-day processing but require cold-chain handling. Heparin tubes should be avoided due to irreversible PCR inhibition.
How does magnetic bead-based extraction compare to filter plate methods for blood RNA?
Magnetic bead methods offer better automation compatibility, lower clog risk, and more consistent inhibitor removal than glass fiber filter plates — processing 96 samples in approximately 17–25 minutes. Filter plates carry higher clog risk when cell lysates are incompletely mixed or sample load exceeds recommended limits.
What RIN score is acceptable for blood-derived RNA in downstream sequencing?
RNA-seq typically requires RIN ≥7, but blood RNA often has lower RIN values due to globin mRNA. DV200 is a more informative quality metric for degraded or complex samples like blood; Illumina recommends DV200 > 70% for high quality, 50-70% for medium, and 30-50% for low quality.
Why does hemoglobin interfere with RNA-based assays and how can it be removed?
Hemoglobin and heme pigments inhibit reverse transcriptase and polymerase enzymes. They are removed through red blood cell lysis, leukocyte isolation, and stringent wash steps during extraction. Visible red/pink eluate is a warning sign of insufficient inhibitor removal.
How do you validate an automated RNA extraction workflow for clinical blood samples?
Validation covers extraction efficiency, repeatability (CV% across replicates), inhibition absence via internal controls, and matrix-specific performance. The workflow must meet applicable regulatory standards — ISO 13485, CE-IVD, or CDSCO IVD guidelines — with documented reports on analytical sensitivity, specificity, and traceability.
Can high-throughput RNA extraction from blood be performed on compact or benchtop instruments?
Yes. Compact automated liquid handlers designed to fit within a biosafety cabinet are well-suited for clinical labs with limited space, provided they are validated for the specific blood RNA extraction chemistry and sample volume in use.


