Introduction
Many clinical diagnostic labs, research institutions, and sequencing centers struggle with inconsistency in nucleic acid extraction—pipetting errors, timing deviations, and technique differences that directly compromise downstream PCR efficiency, NGS library quality, and cell-free DNA yield. Pre-analytical errors account for up to 70% of all mistakes in laboratory diagnostics, and these errors often originate during sample preparation. Even the most advanced analytical instruments cannot correct for poor-quality input material.
Automated sample preparation for nucleic acid extraction uses instruments and software to perform DNA, RNA, and cell-free DNA isolation workflows with minimal manual intervention. It replaces the pipetting, incubation timing, and wash steps operators would otherwise perform by hand.
This matters most in clinical diagnostic labs, research labs, sequencing facilities, and oncology centers, where extraction consistency directly determines the reliability of downstream results.
While automation is frequently recommended for nucleic acid workflows, how it actually works at an operational level—and when it is or isn't the right choice—is rarely explained clearly. This guide covers the underlying mechanics, key instrument categories, validation considerations, and practical criteria for deciding whether automation fits your lab's workflow.
TL;DR
- Automated nucleic acid extraction uses robotics, magnetic bead technology, and pre-programmed liquid handling to isolate DNA/RNA with minimal operator error
- Consistent results depend on a fixed sequence: lysis and binding, washing, and elution — each step building reproducibility into the workflow
- Sample type, bead chemistry, reagent quality, and throughput requirements all shape extraction performance
- Not every lab needs full automation — volume, budget, and workflow complexity should drive the decision
What Is Automated Sample Preparation for Nucleic Acid Extraction?
Automated sample preparation for nucleic acid extraction is a workflow where instruments (liquid handlers or dedicated extraction platforms) execute the full extraction sequence — cell lysis, nucleic acid binding, impurity removal, and elution — without continuous hands-on operator involvement. The result is purified, inhibitor-free nucleic acid at a known concentration and purity ratio, ready for downstream analysis such as PCR, qPCR, NGS, or sequencing.
It's worth understanding how this fits among three distinct workflow types:
- Manual kit-based extraction — uses identical chemistry but requires hands-on pipetting at every step, making it operator-dependent and difficult to scale
- Standalone automated extraction — instruments handle the extraction process independently, then pass purified nucleic acid to a separate analytical platform (PCR, sequencer, etc.)
- Fully integrated platforms — extraction and downstream analysis occur within one closed system, trading flexibility for end-to-end automation
Most labs adopting automation start with standalone extractors. They deliver high-quality nucleic acid templates into existing sequencing or molecular diagnostic workflows without requiring a complete infrastructure overhaul.
Why Labs Are Moving Toward Automated Nucleic Acid Extraction
The Reproducibility Imperative
Manual extraction introduces operator-to-operator and day-to-day variability through pipetting inconsistencies, timing deviations, and technique differences. This variability directly degrades PCR efficiency, sequencing library quality, and cfDNA yield.
One study found that plasma cfDNA extraction efficiencies ranged from 22.9% to 88.1% across specimens, with a coefficient of variance of 28.9%, a spread that illustrates how much manual handling alone can skew results. Automation standardizes these steps, eliminating human variability and delivering consistent results run after run.

Throughput and Turnaround Pressure
Clinical diagnostic labs face rising sample volumes, regulatory expectations for turnaround times, and the reality that a single skilled technician can only run so many manual extractions per shift. Historically, 50 to 60 percent of total molecular testing time in clinical laboratories was spent on manual sample handling and nucleic acid extraction.
Automated platforms compress this timeline substantially, processing dozens of samples in the time it would take to manually prepare a handful.
Cost Reduction Through Efficiency
While the upfront cost of an automated extractor is real, automation reduces cost per test through:
- Reduced hands-on technician time: Automated systems can reduce hands-on time by up to 80-90% compared to manual spin columns
- Fewer failed extractions: Lower error rates mean less rework and fewer wasted reagents
- Higher reagent efficiency: Precise liquid handling eliminates overfilling and waste
- More productive staff deployment: Skilled technicians can focus on higher-value tasks instead of repetitive pipetting
Cambrian Bioworks has published benchmarks showing cost-per-test reductions of up to 56% when moving from manual to automated extraction, driven by these combined efficiency gains.
Contamination and Safety
Manual extraction exposes operators to infectious biofluids, aerosols, and carryover risk across samples. Guidelines recommend waiting five minutes after mixing or vortexing before opening tubes to allow aerosols to settle, highlighting the inherent safety risks in manual workflows. Automated systems with closed processing and built-in decontamination features address both operator safety and cross-sample contamination, which is especially critical in infectious disease and oncology workflows.
Industry Best Practice
In high-throughput clinical and molecular diagnostic labs, automated nucleic acid extraction has moved from aspirational to expected, driven by accreditation bodies, lab quality standards (ISO 15189, ISO 13485), and the practical demands of scaling genomics programs. The global automated nucleic acid extraction market was valued at USD 3.22 billion in 2024 and is projected to reach USD 8.77 billion by 2033, reflecting widespread industry adoption.

How Automated Nucleic Acid Extraction Works: Step-by-Step
Automated nucleic acid extraction takes crude biological samples, processes them through lysis, capture, washing, and elution, and delivers purified nucleic acid — all without the operator pipetting at each stage. Compatible sample types include:
- Blood, plasma, and serum
- Saliva and oral swabs
- Tissue and FFPE sections
- Stool and environmental swabs
Every step is executed by preprogrammed liquid handling movements, making the workflow reproducible across operators and runs.
Technology Approaches
Two dominant technology approaches power automated nucleic acid extraction:
Magnetic bead-based extraction is the most widely adopted method for automation. Under high concentrations of chaotropic salts (such as guanidinium thiocyanate), hydrogen bonds are destabilized, facilitating the binding of nucleic acids to silica-coated paramagnetic particles. An external magnetic field then immobilizes the beads while contaminants are washed away; elution follows by switching to a low-salt, high-pH buffer.
This approach is highly compatible with liquid handling robotics because beads are captured and released magnetically — no centrifugation required.
Spin column-based extraction uses silica membranes in columns but is less amenable to full automation due to its reliance on centrifugation steps. Most purpose-built automated nucleic acid extractors use magnetic bead technology for this reason.
The instrument's software controls timing, volumes, mixing parameters, and temperature. In systems with pre-filled reagent cartridges, manual reagent loading is eliminated entirely — reducing setup error and supporting true walk-away operation.
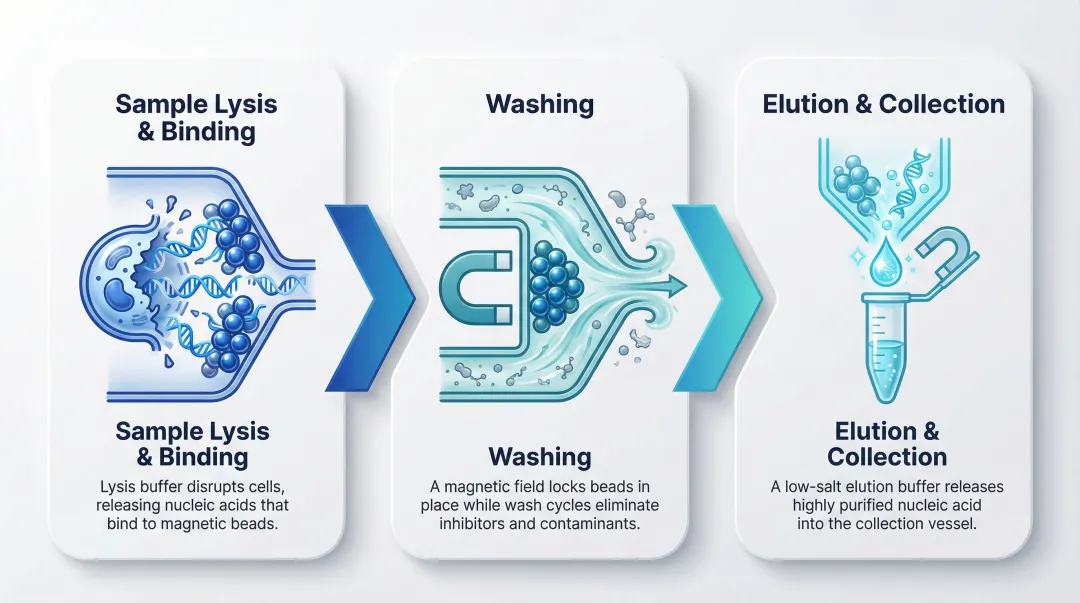
Step 1: Sample Lysis and Nucleic Acid Binding
In this step, a lysis buffer is added to the sample to break open cell membranes and release nucleic acid. Magnetic beads coated with a surface chemistry—such as silica or proprietary ligands—are introduced. Under specific buffer conditions (typically high-salt, low-pH), nucleic acids bind selectively to the bead surface. This step determines binding efficiency and is heavily influenced by sample input type and volume.
Step 2: Washing to Remove Inhibitors and Contaminants
The magnetic field is applied to immobilize the bead-bound nucleic acid while wash buffers are added and removed in sequential cycles. These washes strip away proteins, lipids, cell debris, and PCR inhibitors such as hemoglobin, heparin, or humic acid that would interfere with downstream analysis. Wash stringency and number of wash cycles directly affect purity ratios (A260/A280 and A260/A230). Inadequate washing leaves residual chaotropic salts — even 0.5% guanidinium isothiocyanate contamination can drop the A260/A230 ratio below 0.5, inhibiting downstream enzymatic reactions.
Instruments like Manta by Cambrian Bioworks perform this entire bead-based workflow in approximately 30 minutes with a built-in UVC lamp for inter-run decontamination—addressing both speed and contamination control in clinical environments.
Step 3: Elution and Collection
Once washing is complete, the nucleic acid is released from the beads by applying a low-salt, high-pH elution buffer (or pure water) under controlled temperature and mixing conditions. The purified nucleic acid is transferred to a collection tube or well plate for downstream use. The elution volume is a key variable—lower volumes yield higher concentrations but may reduce total yield. Some automated platforms support elution directly into barcoded tubes, enabling sample traceability critical for clinical lab chain-of-custody requirements.
Key Factors That Affect Automated Extraction Performance
Five variables determine whether automated extraction delivers consistent, reliable results:
Sample type and inhibitor load: Whole blood, plasma, FFPE tissue, saliva, stool, and swabs each present different lysis requirements. Stool contains polysaccharides and bile salts that directly inhibit DNA polymerase, while formalin fixation causes template lesions and fragmentation in FFPE samples. Instrument and chemistry selection must be validated for each specific sample type.
Bead surface chemistry: Silica, carboxyl, streptavidin, and proprietary coatings differ in binding affinity and selectivity. Each is optimized for a specific nucleic acid target — DNA, RNA, or cfDNA. Mismatching bead chemistry to target type is one of the most common causes of low yield.
Reagent condition and storage: Lysis, wash, and elution buffers that are expired or improperly stored degrade extraction efficiency regardless of how well the instrument is configured. Pre-filled cartridge systems reduce this risk by controlling reagent integrity at the point of use.
Instrument format and throughput model: Compact benchtop extractors handling 1–16 samples per run suit labs that process samples on arrival. Higher-throughput liquid handling workstations fit scheduled batch processing workflows. For clinical labs handling urgent samples, batching in automated systems can delay STAT testing — making no-batch-pressure platforms a practical operational advantage.
Open vs. closed system design: Open systems allow third-party reagents and custom protocol development. Closed systems provide validated, locked workflows with tighter quality control. The right choice depends on whether the lab prioritises flexibility or standardisation, and on IVD regulatory requirements.
Common Misconceptions and When Automation May Not Be the Right Fit
Automation Doesn't Compensate for Poor Input Quality
An automated extractor can only work with what it is given. Degraded samples, inhibitor-rich matrices, or improperly stored specimens will produce poor results even on the most sophisticated platform. Automation standardizes the process — consistent execution cannot recover nucleic acid that degradation or contamination has already compromised.
Low-Volume Labs Benefit from Automation Too
Many labs assume automation is only justified for high-volume, high-throughput settings. In reality, even low-volume labs handling 5–20 samples per day benefit from automation through consistency, reduced operator burden, and better documentation. Compact automated extractors are specifically designed for labs that cannot justify large liquid handling workstations.
When Manual Extraction May Still Be Appropriate
Manual extraction may be preferable in several scenarios:
- Extractions requiring custom optimization not yet transferred to an automated protocol
- Settings where capital cost and maintenance overhead outweigh the throughput benefit
- Very low volume settings (1–2 samples per week) where turnaround time is not a constraint
- Forensic and specialized diagnostics: for highly degraded samples such as bone and teeth, phenol-chloroform extraction is still frequently considered the most effective method despite its toxicity and labor intensity
These qualitative thresholds are backed by quantitative data. One study evaluating a magnetic bead-based system found that the automated method became more time-efficient than manual kits only when processing at least 8 samples simultaneously, and became cost-effective after approximately 87 runs of 96 samples — a useful benchmark for any lab weighing the investment against actual throughput.

Conclusion
Automated sample preparation for nucleic acid extraction delivers a standardized, reproducible, walk-away workflow that isolates DNA, RNA, or cfDNA from biological samples with higher consistency, lower error rates, and reduced cost per test compared to manual methods. Magnetic bead chemistry and preprogrammed liquid handling make this possible.
Understanding the process operationally — including what affects performance and where automation isn't appropriate — is what separates a well-implemented system from a poorly matched one.
Labs evaluating automation should assess sample types, throughput needs, and system compatibility before selecting a platform. The right solution delivers measurable improvements in reproducibility, turnaround time, and operator safety when matched to the lab's actual workflow. Platforms like Cambrian Bioworks' Manta and Beluga are designed with exactly this in mind: purpose-built extraction systems that fit clinical and research labs without forcing a compromise on throughput or sample flexibility.
Frequently Asked Questions
What is an automated laboratory system?
An automated laboratory system is an instrument or integrated platform that performs defined analytical or preparative workflows—such as liquid handling, sample extraction, or plate processing—with minimal manual operator input, using robotics, sensors, and software to control each step.
What is the role of automation in clinical laboratory?
Clinical lab automation reduces manual error, increases sample throughput, shortens turnaround time, and supports regulatory compliance. Nucleic acid extraction is one of the most impactful areas to automate given the sensitivity of downstream molecular tests to sample quality.
What are the four types of automation systems?
The four standard categories are pre-analytical automation (sample sorting, labeling, aliquoting), analytical automation (assay execution), post-analytical automation (result reporting, archiving), and total lab automation (TLA), which integrates all three. Nucleic acid extraction sits within pre-analytical or analytical automation depending on platform design.
What is SPE in sample preparation?
Solid-phase extraction (SPE) is a technique where analytes are selectively retained on a solid sorbent and then eluted, primarily used in chromatographic workflows. For nucleic acid extraction, magnetic bead-based methods are the standard choice—they integrate directly with automated liquid handlers and deliver consistent yields across high-throughput runs.
Why is sample preparation important in HPLC?
Sample preparation quality determines whether contaminants and inhibitors are adequately removed before analysis. For nucleic acid assays like PCR and NGS, poor prep leads to inaccurate results and assay failures regardless of how capable the downstream instrument is.


