Introduction
Long-read sequencing platforms like Oxford Nanopore Technologies (ONT) and Pacific Biosciences (PacBio) demand one thing standard NGS workflows rarely deliver: long, intact, highly pure DNA. Next-generation sequencing has become routine for most labs, but high molecular weight (HMW) DNA extraction consistently trips up even experienced teams.
The core problem is mechanical: if your input DNA is fragmented, your reads will be short regardless of the sequencing platform's capabilities. Labs investing in long-read platforms often find their standard extraction protocols destroy the very molecules they need to sequence. The result is fragmented assemblies, missed structural variants, and wasted sequencing capacity.
This article covers what HMW DNA is, why fragment size directly impacts read quality, how the main extraction methods compare, what causes shearing (and how to prevent it), and how to choose the right kit or automated solution.
TLDR
- HMW DNA refers to intact fragments typically exceeding 20–50 kb, with >100 kb preferred for optimal long-read sequencing
- Short-read DNA extraction kits are unsuitable for long-read workflows—they prioritize yield over integrity
- Pipetting force, centrifugation speed, and grinding time are the top mechanical causes of DNA shearing
- Enzymatic lysis with bead- or column-based kits outperforms phenol-chloroform for complex samples
- Automated extraction systems reduce user-introduced shearing and improve reproducibility
What Is High Molecular Weight DNA — and What Size Counts?
HMW DNA refers to exceptionally long, structurally intact DNA molecules spanning hundreds of kilobases to megabases. This distinguishes it from standard genomic DNA (gDNA), which includes fragmented molecules from any extraction method. HMW DNA is defined by both size threshold and structural integrity — not simply by the tissue or sample type it came from.
Size Thresholds That Matter
Fragment analysis commonly uses ≥20 kb as a minimum reference point (percentage of fragments >20 kbp), but most long-read platforms perform optimally with input DNA >40 kb. Applications like optical genome mapping or ultra-long nanopore runs benefit from fragments exceeding 100 kb. For context:
- Standard long-read sequencing: >40 kb preferred
- Ultra-long read workflows: >100 kb ideal
- Optical genome mapping: >150 kb recommended
How HMW DNA Differs from Other DNA Types
Not all DNA types are interchangeable for sequencing workflows. The table below shows how HMW DNA compares to the two most common alternatives:
| DNA Type | Typical Size Range | Primary Use Case |
|---|---|---|
| HMW genomic DNA | >40 kb (up to megabases) | Long-read sequencing, optical genome mapping |
| Cell-free DNA (cfDNA) | ~160–200 bp | Liquid biopsy, prenatal testing |
| Short-read NGS input DNA | 200–400 bp (intentionally fragmented) | Illumina, Ion Torrent platforms |
Neither cfDNA nor fragmented short-read DNA is suitable for long-read workflows, which is why purpose-specific HMW extraction matters.
Why HMW DNA Is the Bottleneck for Long-Read Sequencing
Long-read sequencing platforms generate read lengths proportional to the length of the input DNA molecule that threads through the pore or polymerase. If the input is fragmented, reads are short regardless of the platform's theoretical capability.
Short-Read vs. Long-Read Requirements
Short-read platforms (Illumina, Ion Torrent) are optimized for high throughput with fragmented DNA at 50–300 bp, prioritizing yield and coverage uniformity. They excel at SNP detection and targeted resequencing but struggle with repetitive regions and structural variants.
Long-read platforms require long, contiguous DNA to:
- Span repetitive regions that collapse short-read assemblies
- Resolve structural variants (insertions, deletions, inversions, translocations)
- Support haplotype phasing for distinguishing maternal and paternal alleles
- Enable de novo assembly without a reference genome
- Detect methylation and other epigenetic modifications
Why Conventional Extraction Kits Fail
Kits designed for short-read sequencing often incorporate bead-beating or mechanical cell lysis steps that shear DNA to <10 kb. This maximizes yield and purity but renders DNA unusable for long-read applications. Research by Trigodet et al. found that standard DNeasy PowerSoil with bead beating produced much lower DNA concentrations when modified to reduce shearing. The protocol optimized for yield actively destroys fragment length.
When fragmented input reaches the sequencer, the consequences extend well beyond shorter reads.
Downstream Consequences of Fragmented Input
Shorter reads lead to:
- More fragmented assemblies with hundreds of contigs instead of complete chromosomes
- Higher likelihood of chimeric or composite genomes in metagenomics
- Missed structural variants in oncology applications
- Incomplete haplotype phasing in rare disease genetics
Research by Koren et al. (2013) demonstrated that approximately 7 kb reads can span most repetitive regions. Reads below this threshold significantly compromise assembly quality.
Clinical and Research Applications
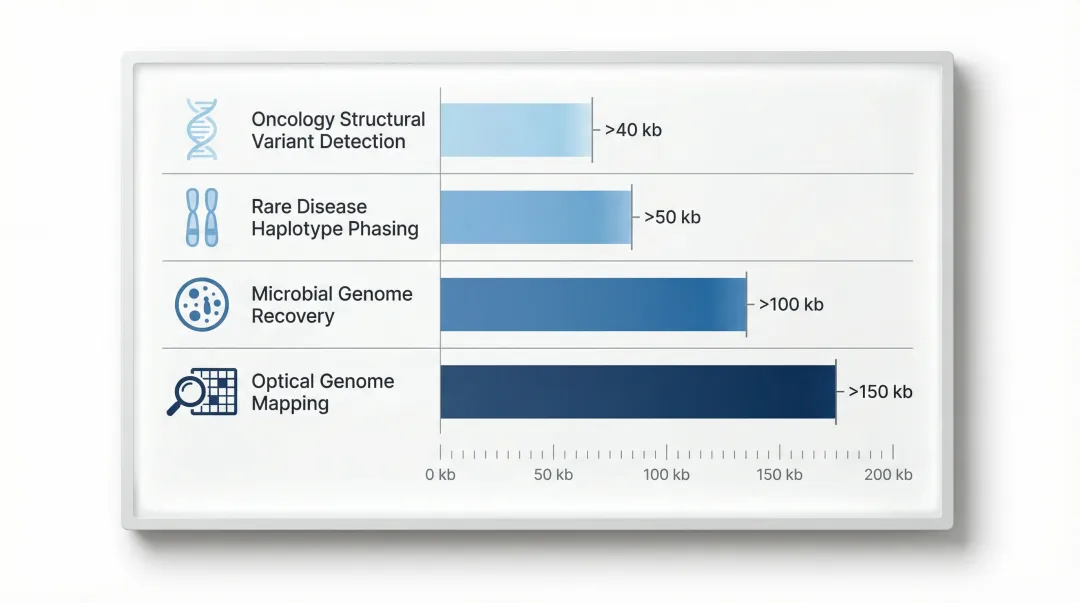
HMW DNA quality is especially critical for:
- Oncology structural variant detection requires >40 kb fragments to span chromosomal breakpoints
- Rare disease haplotype phasing needs >50 kb to connect distant variants across a locus
- Complete microbial genome recovery benefits from >100 kb for single-contig assemblies
- Optical genome mapping demands >150 kb fragments for reliable alignment
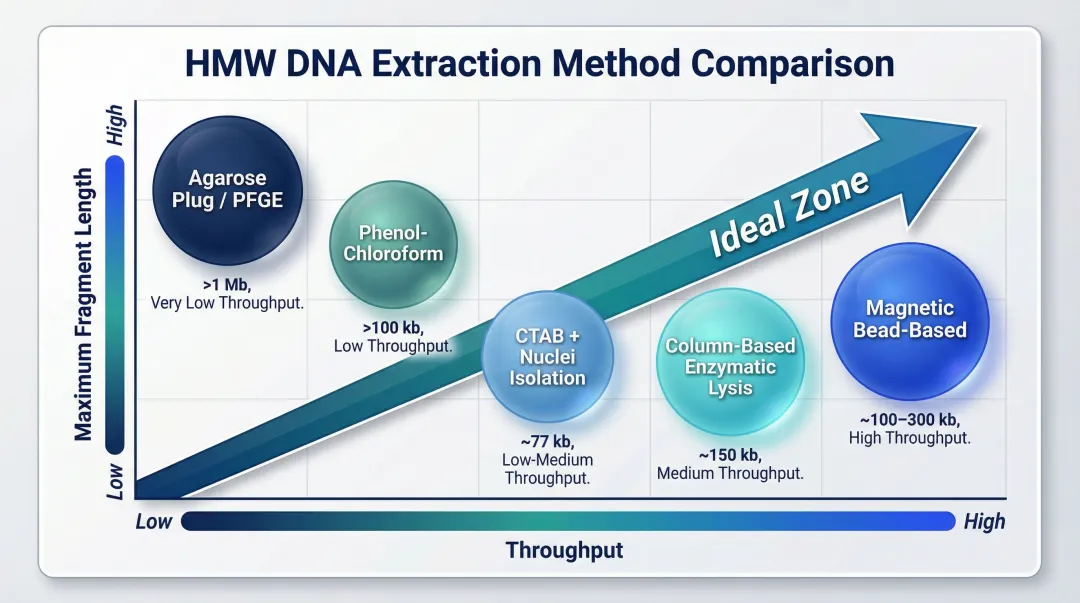
HMW DNA Extraction Methods: A Practical Comparison
Four main approaches dominate research and clinical labs: phenol-chloroform extraction, column-based kits with enzymatic lysis, magnetic bead-based purification, and agarose plug encasement. Choosing between them comes down to your fragment length targets, sample type, and throughput requirements.
Phenol-Chloroform Extraction
What it delivers: The longest individual DNA fragments (>100 kb reported using the Sambrook method)
Significant drawbacks:
- Hazardous reagents requiring fume hoods and specialized disposal
- High eukaryotic host DNA contamination in mixed samples (up to 81% human reads in oral metagenome studies)
- Very low proportion of medium-length reads useful for assembly
- Gram-negative bias due to lack of bacterial-specific lytic enzymes
While phenol-chloroform can produce the longest fragments, practical performance in complex samples is poor. The method excels only in controlled contexts with pure cell cultures.
Column-Based Kits with Enzymatic Lysis
Best representatives: Qiagen Genomic Tip series with enzyme supplementation
Replacing mechanical bead beating with enzymatic lysis (lysozyme, mutanolysin, proteinase K cocktails) consistently delivers the best balance of yield, purity, and read-size distribution. The Trigodet et al. study found that the Genomic Tip (GT) method yielded the highest DNA concentration (mean ~110 µg/mL), best purity (A260/280 ~1.8), and the most circular assembled elements in metagenomics applications.
Key advantages:
- Gentle gravity-flow purification minimizes shearing
- Fragments up to 150 kb routinely achieved
- Yields 20–500 µg depending on column size
- Compatible with diverse sample types
Limitations:
- Manual protocol requires careful technique
- Lower throughput than automated methods
- Not easily scalable for batched processing
Magnetic Bead-Based Extraction
Magnetic beads selectively bind DNA under specific salt and buffer conditions. The key advantage is gentle handling: no spin columns, no harsh centrifugation, and full compatibility with automation.
Bead-based methods can be optimized for HMW enrichment by tuning bead ratios. Higher bead:sample ratios favor shorter fragments; lower ratios can preferentially capture longer molecules. This selectivity is the core principle behind size-selective magnetic bead workflows.
Performance data from vendor comparisons (Thermo Fisher, 2025) show the MagMAX HMW DNA Kit achieves 80.70% of fragments >40 kb and 76.73% >100 kb from whole blood, though vendor-supplied benchmarks should be independently validated.
Key advantages:
- Automation-ready workflows
- Consistent run-to-run reproducibility
- Scalable throughput (1–96 samples)
- Compatible with magnetic bead processors like KingFisher systems
CTAB and Nuclei Isolation Methods (Plant and Complex Tissue)
For plant genomics, CTAB-based protocols combined with nuclei isolation reduce organellar DNA contamination. The nuclei are isolated first, followed by CTAB lysis and chloroform-isoamyl alcohol purification.
Performance data: Research shows this combined protocol produces DNA fragments approximately 5× longer on average (peak ~77.88 kbp, ~67.5% of fragments >20 kb) versus commercial kits (~18.65 kbp average, ~36% >20 kb).
Limitations:
- Requires experienced technicians
- Multiple manual steps increase variability
- Less amenable to automation
- Time-intensive protocols (several hours)
Agarose Encasement (PFGE Plugs)
Embedding cells in agarose prevents shearing entirely and can yield DNA >1 Mb. Cells are lysed within the plug, then digested with agarase to release ultra-long DNA.
Critical caveat: More is not always better. Very long fragments (>500 kb) may become trapped in sequencing pores or lost during library preparation without corresponding protocol optimization.
Practical limitations:
- Very low throughput (typically 6–12 samples per run)
- Labor-intensive multi-day protocols
- Difficult to standardize across operators
Each method occupies a different point on the speed-vs-fragment-length curve. The sections below cover how kits and automation systems map onto these methods — and where purpose-built HMW solutions close the remaining gaps.
Key Factors That Cause DNA Shearing — and How to Avoid Them
Key Factors That Cause DNA Shearing and How to Avoid Them
Pipetting Mechanics
High-speed or frequent pipetting creates velocity gradients that physically shear long DNA strands. Empirical data shows that 200 pumps at maximum speed reduced fragments >50 kb from 17% to 13.4% and shifted the peak from 76.46 kbp to 49.93 kbp.
To protect fragment integrity during liquid handling:
- Use wide-bore pipette tips (1000 µL)
- Pipette slowly and gently
- Limit pipetting steps to fewer than 20 where possible
- Use end-over-end rotation instead of vortexing for mixing
- Allow DNA to flow by gravity rather than forcing aspiration
Centrifugation Speed
High centrifugal forces cause DNA molecules to collide and fragment. Data demonstrates that 5000 × g reduced >50-kb fragments from 39.7% to 30.0% and shifted the peak from 146.94 kbp to 88.73 kbp versus 3000 × g.
Keep centrifugation at or below 3000 × g for all HMW DNA workflows. High-speed spins at any step involving intact HMW molecules risk irreversible fragment loss.
Mechanical Lysis and Grinding
Excessive grinding — especially in plant tissue protocols — and bead-beating can improve yield, but at a real cost to fragment length. Additional grinding reduced fragment length distributions and diminished the proportion of >50-kb fragments.
For samples requiring disruption:
- Replace mechanical homogenization with enzymatic digestion using proteinase K wherever possible
- For plant material, limit grinding to 1 minute — extra liquid nitrogen beyond that is counterproductive
- Avoid bead-beating protocols for HMW applications
- Use enzymatic lysis cocktails tailored to sample type

Sample Handling, Storage, and Freeze-Thaw
HMW DNA is vulnerable long after extraction. Residual nucleases remain active at room temperature, and each freeze-thaw cycle introduces mechanical stress that shortens fragments. Improper storage temperature and prolonged incubation compound the damage.
Protect sample integrity post-extraction with these precautions:
- Store eluted HMW DNA at 4°C short term, −20°C for longer storage
- Limit freeze-thaw cycles to one
- Use LoBind tubes to reduce adsorption of long DNA to tube walls
- Keep samples on ice during processing
- Include RNase A and proteinase K treatments to clear RNA and protein contamination
Commercial Kits and Automated Solutions for HMW DNA Extraction
Kit Categories and Key Differentiators
The commercial HMW DNA kit landscape includes magnetic bead-based kits (MagMAX HMW DNA Kit, Wizard HMW Extraction Kit), column-based kits with enzymatic lysis options (Qiagen Genomic Tip series), and specialized technologies.
Key differentiators:
- Compatible sample types
- Minimum input requirements
- Maximum fragment size achievable
- Automation compatibility
- Regulatory certification (IVD vs. RUO)
Comparison of Leading Commercial Kits
| Kit Name | Sample Types | Max Fragment Size | Typical Yield | Automation Compatible | Purity Targets |
|---|---|---|---|---|---|
| Thermo Fisher MagMAX HMW | Blood, cells, tissues, saliva, buccal swabs | 100–300 kb | Variable by input | Yes (KingFisher) | A260/280: ~1.8; A260/230: ≥2.0 |
| Promega Wizard HMW | Blood, cells, plant tissue, bacteria | Up to 500 kb | 12.5 µg from 300 µL blood | Manual protocol | A260/280: ~1.8; A260/230: ≥2.0 |
| Qiagen Genomic-tip | Bacteria, blood, cells, tissue, yeast, plants | Up to 150 kb | 20–500 µg (size dependent) | No | A260/280: ~1.8; A260/230: ≥2.0 |
What to Evaluate When Choosing a Kit
Minimum DNA yield requirements: Most long-read protocols need ≥1–1.5 µg input
Purity targets: Both A260/A280 and A260/A230 ratios should exceed 1.8, with A260/A230 ≥2.0 preferred to prevent enzymatic inhibition during library preparation. Oxford Nanopore Technologies recommends A260/A230 ratios of 2.0–2.2 for optimal sequencing performance.
Fragment size distribution: Look for >40 kb peak, ideally median well above 20 kb
Platform validation: Verify the kit has been tested on your specific sequencing platform (ONT vs. PacBio)
Automation as a Solution to User-Introduced Variability
One major source of HMW DNA degradation is inconsistent manual technique—variation in pipetting speed, mixing force, drying time, and digestion temperature between operators. Automated liquid handlers and magnetic bead processors eliminate these variables, delivering consistent fragment size distributions run-to-run.
Cambrian Bioworks' CamSelect Long™ Bead Technology, a patented magnetic bead platform engineered specifically for HMW DNA enrichment, addresses a fundamental constraint in long-read sequencing: finite flowcell capacity. Short DNA fragments occupy pores unproductively and accelerate pore attrition, wasting sequencing capacity. CamSelect Long™ applies tunable size-selective enrichment with cutoff options at under 2 kb, 5 kb, 7 kb, and 10 kb, selectively depleting short fragments while retaining longer molecules.
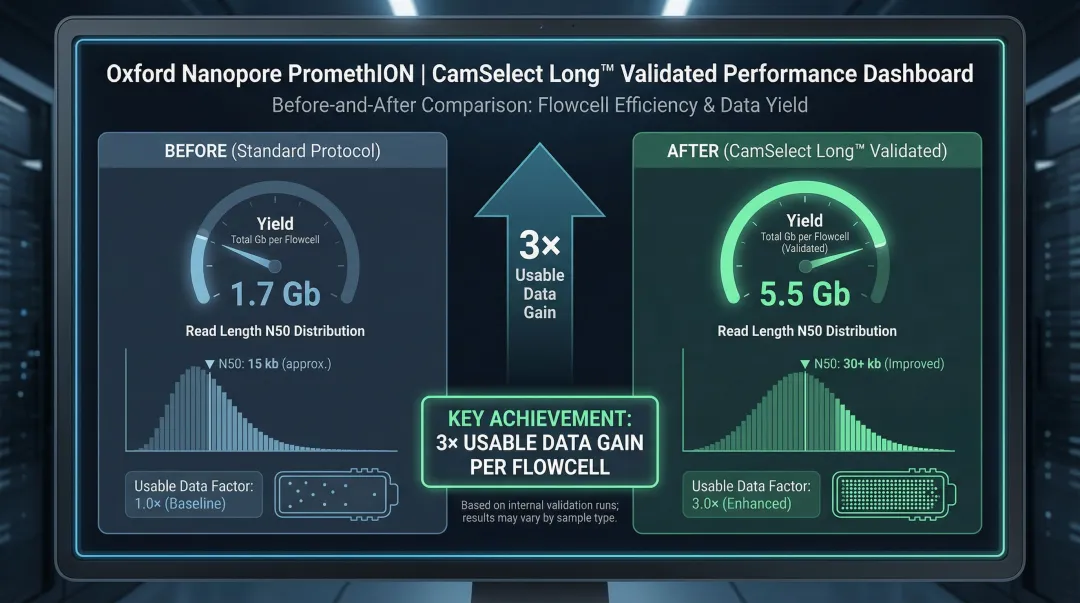
Validated performance on Oxford Nanopore PromethION platforms demonstrates measurable gains:
- 3× increase in usable data per flowcell
- 223.5% improvement in total yield (1.7 Gb → 5.5 Gb)
- 56.4% increase in read length N50 (4,745 bp → 7,419 bp)
- Improved long-read fraction and enhanced pore retention

Combined with Cambrian Bioworks' **Beluga automated liquid handler**, labs achieve standardized HMW DNA extraction at scale with no batching pressure. The Beluga platform enforces consistent pipetting speeds, controlled aspiration rates, and reproducible mixing protocols, removing the operator-dependent variables that degrade fragment integrity in manual workflows.
Measuring HMW DNA Quality Before Sequencing
Quantification Methods
Two complementary methods give you the full quality picture before sequencing:
| Method | What It Measures | Key Limitation |
|---|---|---|
| NanoDrop (spectrophotometric) | Total nucleic acid; A260/A280 and A260/A230 ratios | Overestimates concentration when RNA or contaminants are present |
| Qubit (fluorometric) | Double-stranded DNA specifically | Does not detect contamination ratios |
Both ratios should fall between 1.8–2.2 for DNA free of protein, phenol, and carbohydrate contamination. Run both instruments together — NanoDrop flags contamination, Qubit confirms true DNA yield.
Fragment Size Assessment Tools
Standard gel electrophoresis and TapeStation are insufficient for HMW DNA—they cannot accurately separate fragments >60 kbp.
For fragments above 60 kbp, three sizing options are available — with meaningful differences in accuracy and workflow:
- Femto Pulse (Agilent): Automated pulsed-field electrophoresis; separates DNA up to 165 kbp, making it the go-to tool for fragments above 60 kbp in most long-read sequencing workflows
- Pippin Pulse / PFGE: Valid alternatives, but require careful DNA loading normalization — the quantity loaded affects apparent band position
- Bioanalyzer High Sensitivity DNA: Optimized for 35–10,380 bp only; not suitable for HMW sizing above ~10 kb
Interpreting Fragment Size Data
Good HMW DNA on a Femto Pulse trace shows:
- A peak well above 20 kbp (ideally >40 kbp for most long-read applications, >100 kbp for ultra-long reads)
- High percentage of fragments in the HMW range (>20 kbp)
- Minimal low-molecular-weight smearing
Troubleshooting guide:
- Peak <20 kbp: Extraction conditions need revision—check pipetting technique, centrifugation speed, and lysis method
- A260/A230 <1.8: Chemical contamination present (phenol, carbohydrates, guanidine)
- A260/A280 >2.0: RNA contamination likely—add RNase A treatment
Frequently Asked Questions
What is high molecular weight DNA extraction?
HMW DNA extraction is a specialized isolation process designed to recover long, intact DNA molecules (typically >20–100 kb) from biological samples while minimizing shearing. Unlike standard DNA extraction that prioritizes yield, HMW extraction prioritizes fragment length integrity essential for long-read sequencing applications.
What is considered high molecular weight DNA?
HMW DNA generally refers to DNA fragments exceeding 20–50 kb, with many long-read sequencing applications requiring a predominance of fragments >40 kb. Ultra-long read workflows benefit from DNA peaking above 100 kb, and optical genome mapping applications prefer fragments >150 kb.
What is a good A260/A280 ratio for DNA?
An A260/A280 ratio of approximately 1.8 indicates pure double-stranded DNA. Values significantly below 1.8 suggest protein or phenol contamination, while values above 2.0 indicate RNA contamination.
What is the size range for the Bioanalyzer High Sensitivity DNA assay?
The Bioanalyzer High Sensitivity DNA assay is optimized for fragments in the range of approximately 35 bp to 10,380 bp, making it suitable for library QC but not for characterizing HMW DNA fragments above approximately 10 kb. For true HMW fragment sizing, the Femto Pulse system is the recommended alternative.
What are alternatives to the Bioanalyzer for DNA sizing and quantification?
Several tools outperform the Bioanalyzer for HMW applications:
- Femto Pulse (pulsed-field capillary electrophoresis) resolves DNA up to 165 kbp — the preferred tool for true HMW sizing
- Pulsed-field gel electrophoresis (PFGE) separates large DNA fragments across a wide size range
- Qubit fluorometry delivers accurate dsDNA quantification, unaffected by free nucleotides or degraded fragments
- NanoDrop spectrophotometry provides A260/A280 and A260/A230 purity ratios
A combination of Qubit and Femto Pulse is current best practice for HMW DNA quality assessment.


