Introduction
Library preparation quality is directly determined by RNA input quality. Blood and FFPE are two of the most clinically valuable yet technically demanding sample types, each presenting different degradation and contamination profiles. Blood RNA degrades rapidly post-collection due to RNase activity, while FFPE RNA is fragmented by formalin chemistry over months to years.
According to a 2014 study published in BMC Genomics, RNA degradation distorts read coverage and transcript quantification, skewing differential expression calls and reducing library complexity. This makes pre-analytical quality control— not just library prep kit selection — a foundational step of reliable RNA-Seq data.
What follows covers the distinct RNA quality thresholds for blood and FFPE samples before library prep begins, the metrics that actually matter for each, and practical strategies to salvage challenging inputs.
TLDR
- Blood RNA: Aim for RIN ≥7, A260/280 of 1.8–2.1, and 500 ng–1 µg input— globin contamination (70%+ of mRNA) and hemolysis are the top quality threats
- FFPE RNA: RIN is unreliable—use DV200 instead; DV200 >66% predicts library success, 30–50% is marginal
- Sample collection (tube type, fixation time, storage) drives RNA integrity before extraction begins
- Automated extraction reduces inter-sample variability for consistent library prep inputs
Why RNA Input Quality Is the Foundation of Successful RNA-Seq
Poor RNA input triggers a cascading quality failure: each downstream step compounds the damage from the last.
- Inefficient reverse transcription produces biased or truncated cDNA fragments
- Low library complexity follows, reducing the representational depth of your sequencing run
- The result: noisy, unreliable data with skewed differential expression calls
Library prep kits cannot rescue severely degraded RNA. They can compensate for moderate degradation, but starting material quality sets the ceiling on what any kit can achieve. FFPE-optimized kits using random priming and ligation-based construction still require a minimum DV200 threshold — below 30%, library failure risk is high.
That threshold varies by sample type, because blood and FFPE samples degrade through entirely different mechanisms. Blood RNA deteriorates rapidly post-collection due to RNase activity, while FFPE RNA is fragmented by formalin chemistry over months to years. Applying a single quality cutoff across both sample types sets up workflows — and study conclusions — for failure.
Blood Samples: RNA Quality Challenges and Input Requirements
Collection Tube and Stabilization: The First Determinant
PAXgene tubes preserve transcriptomes in situ within minutes of collection. They contain proprietary reagents that lyse cells and stabilize intracellular RNA for up to 3 days at 18–25°C. RNA extracted from PAXgene tubes maintains high integrity (RIN >8) even after 7 to 11 years of cryopreservation at -80°C.
EDTA tubes require immediate processing or cold-chain preservation. Stability at 4°C is maintained for up to 168 hours, but at room temperature (25°C) only for 72 hours — making logistics a real constraint for multicenter studies.
For frozen EDTA blood, a thaw-into-lysis protocol — adding RNA stabilizing buffer directly during the thaw — improves RIN to 7.3–8.0 and yields five times more RNA compared to standard extraction. This step prevents reactivated RNases from degrading RNA before extraction begins.
Hemolysis: A Critical Confound
Hemolyzed samples release hemoglobin and cell-free hemoglobin mRNA that inhibit reverse transcriptase. Severe hemolysis (free hemoglobin >4 g/L) significantly decreases library concentration, increases duplication rates, and elevates test failure risk.
Use a hemolysis index threshold to flag samples before committing to library prep:
- Slight or moderate hemolysis (≤4 g/L): Generally acceptable for most protocols
- Severe hemolysis (>4 g/L): Skews expression profiles toward erythroid transcripts, reduces library complexity, and risks assay failure
- Practical rule: Flag at the pre-analytical stage — downstream rescue options are limited once library prep begins
Globin mRNA Dominance
In whole blood, α- and β-globin transcripts from erythroid precursors account for 70% to over 80% of the total mRNA. If not depleted, these highly abundant transcripts dominate sequencing output, masking biologically informative transcripts and reducing sensitivity for low-abundance genes.
Minimum Quality Thresholds for Blood RNA Library Prep
RIN ≥7 for standard whole-transcriptome kits
Lower RIN (≥6) may be tolerated by some ultra-low input or ribo-depletion-based protocols, but expect increased 3' bias and reduced library complexity.
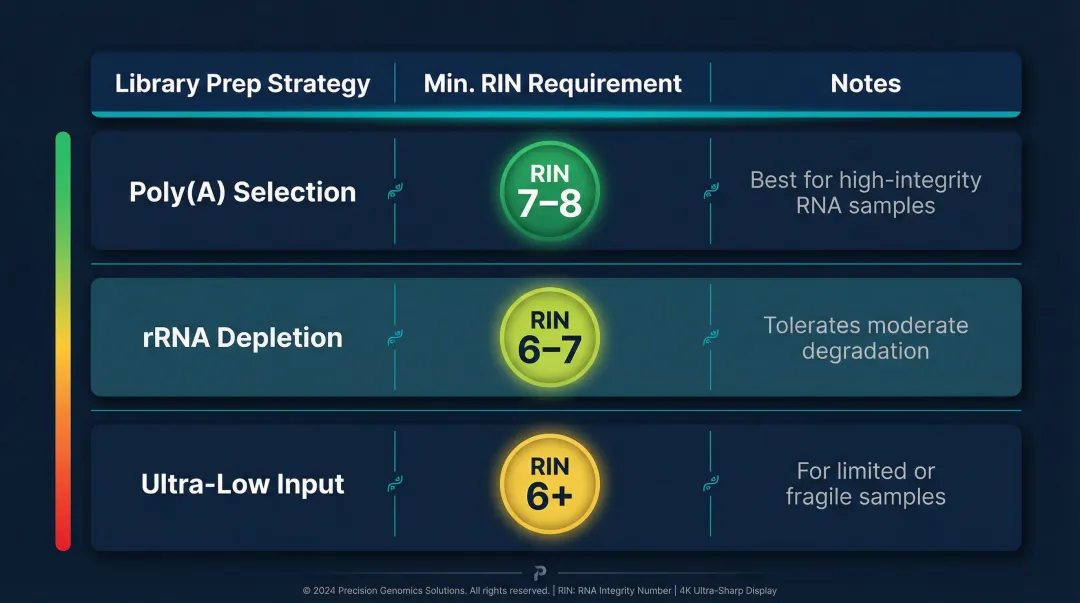
| Library Prep Strategy | Minimum RIN | Notes |
|---|---|---|
| Poly(A) selection | 7–8 | Requires intact poly(A) tails |
| rRNA depletion | 6–7 | More tolerant of degradation |
| Ultra-low input | 6+ | Lower input amplifies quality issues |
Input Quantity Guidelines
Standard blood RNA-Seq library prep protocols typically require 500 ng–1 µg of total RNA. Low-input protocols can work from as little as 10–100 ng, but lower input amplifies any pre-existing quality issues and increases PCR duplication rates.
FFPE Samples: RNA Quality Challenges and Input Requirements
The Biochemistry of Formalin-Induced RNA Damage
Formalin fixation induces two destructive mechanisms:
- Crosslinking: Formaldehyde adds mono-methylol groups to RNA bases and creates methylene bridges that crosslink RNA to proteins
- Hydrolytic degradation: Phosphodiester bonds fragment the RNA backbone into sub-200 nt fragments during paraffin embedding and storage
Heating in formalin-free buffer reverses most methylol crosslinks, but the fragmentation is permanent.
Why RIN Fails for FFPE
The RIN algorithm was designed for intact RNA with two clear ribosomal peaks (28S and 18S). FFPE RNA produces a degraded low-molecular-weight smear that yields artificially low or undefined RIN scores.
DV200 is the accepted standard. DV200 is the percentage of RNA fragments larger than 200 nucleotides. Receiver operating characteristic (ROC) curve analyses reveal that a DV200 cutoff of 66.1% provides greater sensitivity (92%) and specificity (100%) than RIN for predicting efficient library production.
DV200 thresholds:
- >66%: High quality, optimal for library prep
- 50–66%: Medium quality, acceptable with higher input
- 30–50%: Marginal quality, high risk for library failure
- <30%: Not recommended
Impact of Fixation Duration and Storage Age
Longer formalin fixation times (>24–48 hours) and samples archived for multiple years produce markedly lower DV200 values. Tissues fixed within the 12–24 hour window yield RNA with significantly higher DV200 compared to those over-fixed at 48 hours or beyond. Prolonged storage of FFPE blocks at room temperature leads to progressive DV200 decline.
Library Prep Strategy Adapts to FFPE RNA Fragmentation
These DV200 tiers directly shape which library prep approach is viable. Poly(A) selection is not appropriate for FFPE: formalin fixation frequently degrades poly(A) tails, so oligo(dT) capture produces severe 3' bias and significant loss of transcript diversity. rRNA depletion or targeted capture are the practical alternatives, using ligation-based approaches with random priming to recover even heavily fragmented transcripts.
| Library Prep Strategy | Exonic Mapping Rate | Performance on Degraded RNA |
|---|---|---|
| TruSeq RNA Access (Capture) | 81–84% | High concordance; robust down to DV200 30% |
| TruSeq Stranded Total (rRNA Depletion) | 17–41% | High intronic/intergenic reads; requires higher input |
| TaKaRa (rRNA Depletion) | 48–64% | Tolerates ultra-low input (10 ng) but leaves high rRNA residual |
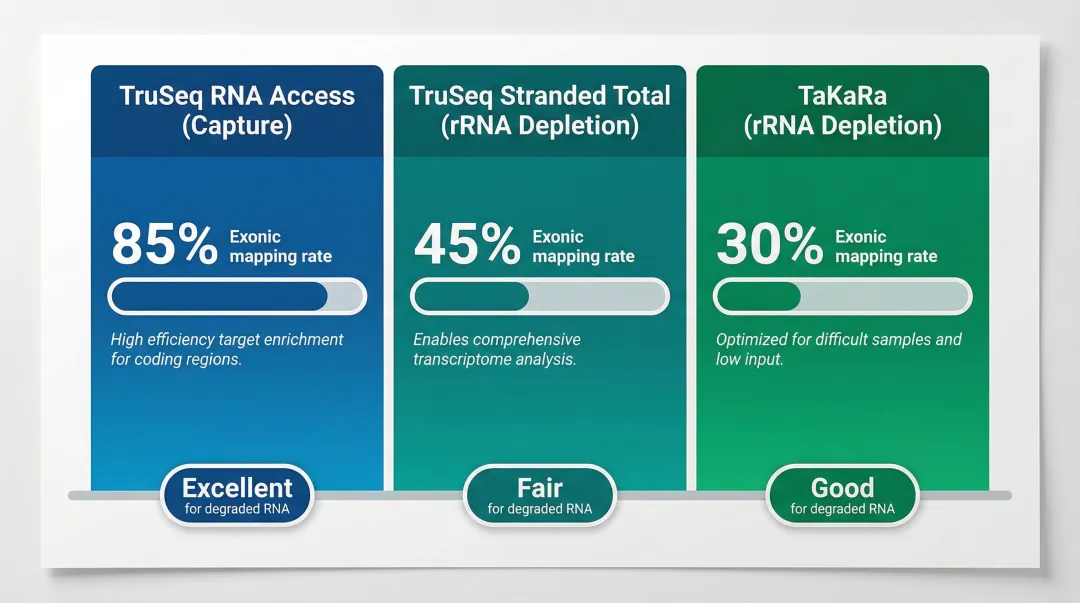
Exome capture yields >80% exonic reads versus 17–41% for rRNA depletion in FFPE, making targeted capture the preferred approach for gene-focused studies.
Input Quantity for FFPE
Typically 100 ng–1 µg depending on kit and DV200. Higher input partially compensates for lower-quality RNA but increases rRNA background. Advanced low-input FFPE protocols can process as little as 10–50 ng from needle biopsy or laser capture microdissection (LCM) material.
How to Assess RNA Quality Before Library Prep
Spectrophotometric Assessment (NanoDrop)
Useful for: Estimating concentration and purity via A260/280 (target 1.8–2.1 for pure RNA) and A260/230 (target >1.8) to flag protein, phenol, or salt contamination.
Key limitation: NanoDrop cannot detect degradation, only contamination and concentration. UV spectrophotometry frequently overestimates RNA concentration compared to fluorometric methods because it cannot distinguish between intact RNA, free nucleotides, or degraded nucleic acids.
Fluorometric Quantification (Qubit)
Preferred method for accurate RNA concentration measurement. Qubit uses intercalating dyes specific to RNA and is not affected by free nucleotides, salts, or protein. Always use Qubit or equivalent for final input concentration decision before library prep.
Capillary Electrophoresis (Bioanalyzer/TapeStation)
Gold standard for RNA integrity assessment. Generates RIN for intact samples and fragment-size electropherograms. For FFPE, use the TapeStation RNA ScreenTape assay or equivalent to calculate DV200 directly.
How to read an electropherogram:
- Blood/intact RNA: Look for two clear peaks at 28S and 18S. RIN ≥7 indicates minimal degradation
- FFPE RNA: Look for a degraded smear with no clear ribosomal peaks. Calculate DV200 (percentage of fragments >200 nt) using software tools
Automated Extraction for Consistent, Reproducible Input Quality
QC results downstream are only as reliable as the extraction step upstream. Standardizing extraction directly controls one of the most variable inputs in the workflow — manual extraction introduces operator-dependent variability in both RNA integrity and yield.
Cambrian Bioworks' automated nucleic acid extraction platforms (Manta and Beluga) deliver reproducible RNA extraction from challenging clinical samples, including blood and FFPE tissue. The Manta platform is purpose-built for clinical FFPE workflows:
- Processes 1–32 samples per run with minimal hands-on time
- Completes FFPE RNA extraction in approximately 2.5 hours
- Requires only four 10 µm FFPE curls per sample
- Consistently yields approximately 5,000 ng of RNA with DV200 values >70%

Strategies to Improve Challenging RNA Inputs Before Library Prep
Globin Depletion for Blood Samples
For whole blood RNA, globin mRNA removal (using bead-based capture or enzymatic methods) raises the proportion of informative transcripts sequenced.
Trade-off: Some depletion protocols can reduce overall yield and must be performed on high-enough quality starting RNA (RIN ≥6).
| Depletion Method | Residual Globin % | Coverage | Limitations |
|---|---|---|---|
| Probe hybridization (GLOBINclear) | <1% | Uniform gene body | Requires high input (1 µg); slight RIN drop |
| RNase-H enzymatic (Ribo-Zero Plus) | 3.2% | Skewed to 3' region | Enzymatic degradation alters RNA quality |
A 2021 comparative study (PMC7733259) found that probe-hybridization methods detect more genes and transcripts without the 3' region bias introduced by RNase-H enzymatic depletion.
DNase Treatment to Eliminate gDNA Contamination
Residual genomic DNA co-purified with RNA can ligate to adapters and generate spurious reads during sequencing. Blood cells contain significantly more DNA than RNA, making genomic DNA (gDNA) carry-over highly likely during extraction.
DNase I treatment — either on-column or in-solution — should be a standard step, particularly for blood-derived RNA where buffy coat preparation can co-extract leukocyte genomic DNA. In-solution DNase I digestion consistently outperforms on-column methods in both efficiency and completeness of gDNA removal.
rRNA Depletion as the Preferred Enrichment Strategy for FFPE
Unlike poly(A) selection, rRNA depletion works on fragmented RNA and captures the full transcriptome including non-polyadenylated transcripts. When exonic mapping is the goal, however, the two strategies diverge sharply:
- rRNA depletion: 17–41% exonic mapping efficiency; broader transcriptome coverage including lncRNAs
- Targeted capture: 81–84% exonic mapping efficiency; preferred when gene-focused expression data is the priority
Selecting the right pre-treatment strategy for your sample type is the foundation that determines whether library prep succeeds or fails — the steps outlined above directly shape yield, quality, and downstream sequencing performance.
Frequently Asked Questions
What does library preparation mean in sequencing?
Library preparation converts extracted RNA (or DNA) into a sequencing-ready collection of adapter-flanked cDNA fragments. The process covers reverse transcription, fragmentation, adapter ligation, and amplification — producing fragments the NGS platform can bind, cluster, and read.
What is the minimum RNA input required for RNA-Seq library preparation from FFPE samples?
Most standard FFPE library prep kits require 100 ng–1 µg, while specialised low-input protocols can work from as little as 10–50 ng. Higher inputs are recommended when DV200 is below 50% to compensate for reduced RNA quality.
How does hemolysis in blood samples affect RNA-Seq library quality?
Haemolysed samples release haemoglobin and cell-free haemoglobin mRNA that inhibit reverse transcriptase. This skews expression profiles toward erythroid transcripts and reduces library complexity. Severe haemolysis (>4 g/L free Hb) increases duplication rates and decreases library concentration.
What is DV200 and why is it used for FFPE RNA quality assessment?
DV200 is the percentage of RNA fragments larger than 200 nucleotides. It is more informative than RIN for FFPE samples because it directly quantifies recoverable fragment length. RIN relies on intact ribosomal RNA peaks, which are absent in degraded FFPE material.
Can degraded RNA from FFPE be used for RNA-Seq library preparation?
FFPE RNA with DV200 between 30–50% can yield libraries using FFPE-optimised kits with random priming and rRNA depletion. Library complexity and gene detection rates will be reduced compared to higher-quality inputs (DV200 >66%), so managing expectations around output depth is important.
What is the difference in library prep approach between whole blood and FFPE samples?
Blood RNA prep typically prioritises globin depletion and may use poly(A) selection if RIN is high, whereas FFPE prep requires rRNA depletion or capture-based methods and FFPE-optimised fragmentation-aware kits due to pre-existing RNA fragmentation.


