Introduction
Formalin-fixed paraffin-embedded (FFPE) tissue blocks represent one of the most valuable yet underutilised resources in modern genomics. Hospital archives hold decades of FFPE samples paired with clinical annotations on patient outcomes, treatment responses, and pathology—data that could yield actionable insights into disease progression. Yet many sequencing labs hesitate to work with FFPE material at all.
The hesitation is understandable. Uncertainty around RNA quality assessment, extraction protocol selection, and kit performance keeps archival tissue sitting on shelves rather than generating data.
Three critical pain points prevent labs from confidently incorporating FFPE tissue into RNA-seq workflows:
- Deparaffinisation and crosslink reversal steps that vary significantly between protocols — with no clear consensus on best practice
- Commercial kits that differ in yield, purity, and suitability depending on input quantity, tissue age, and throughput requirements
- RIN thresholds borrowed from fresh/frozen workflows that cause labs to discard FFPE samples that would produce usable sequencing data
This guide works through each challenge with research-backed recommendations. You'll find extraction protocols ranked by evidence, a structured kit comparison across common input conditions, and a practical walkthrough of RNA quality metrics — including why DV200 and electropherogram curve-shape analysis are better predictors of sequencing success than RIN scores alone.
TLDR
- FFPE RNA is fragmented and chemically modified; RIN scores of 1–3 are normal and do not predict failure—use DV200 instead
- DV200 ≥30% is workable; for borderline samples, the electropherogram curve shape near 100 bp indicates usable fragment yield
- Qiagen AllPrep DNA/RNA FFPE supports dual DNA/RNA extraction; TaKaRa SMARTer Stranded Total RNA works from inputs as low as 5 ng
- Always quantify by fluorescence (Qubit)—NanoDrop overestimates concentration roughly 2-fold
- rRNA depletion is mandatory—poly-A selection fails on FFPE because degraded tails are too short to capture
Why FFPE RNA-Seq Is Technically Challenging
The Chemistry Behind Formalin Fixation
Formaldehyde fixation fundamentally alters RNA structure through methylene bridge formation between amino groups on nucleotides (A, C, G, U). This crosslinking reaction causes:
- Strand fragmentation: RNA molecules break into fragments typically shorter than 200 nucleotides
- Base modification: Chemical adducts form on nucleotide bases, altering their structure
- Protein-RNA crosslinking: RNA becomes covalently bound to proteins, complicating purification
Unlike degradation in fresh or frozen samples—which results from RNase activity—formalin-induced damage cannot be fully reversed. The chemistry is permanent, leaving fragments that behave differently from intact RNA in downstream workflows.
Downstream Consequences for Library Preparation
Three major technical barriers emerge from formalin damage:
- Poly-A capture fails: Poly-A tails degrade preferentially during fixation. Oligo-dT protocols lose the majority of transcripts, making total RNA with rRNA depletion the required approach.
- RIN values are unreliable: The RIN algorithm expects intact 18S and 28S ribosomal bands. FFPE samples universally lack these, producing RIN scores of 1–3 even when samples are sequencing-ready.
- GC-rich regions are systematically biased: Disproportionate crosslinking causes high-GC genes to appear underrepresented in expression profiles, requiring computational correction during analysis.
Fixation Quality Matters More Than Block Age
Research demonstrates no statistically significant correlation between block storage time—even up to 20 years—and RNA yield per unit tissue area. Age alone doesn't predict failure — fixation duration does:
- Over-fixation (formalin exposure beyond 48 hours) significantly worsens crosslinking and increases fragmentation
- Under-fixation produces poor tissue morphology and inconsistent RNA recovery
- Optimal fixation: 12–24 hours in neutral buffered formalin
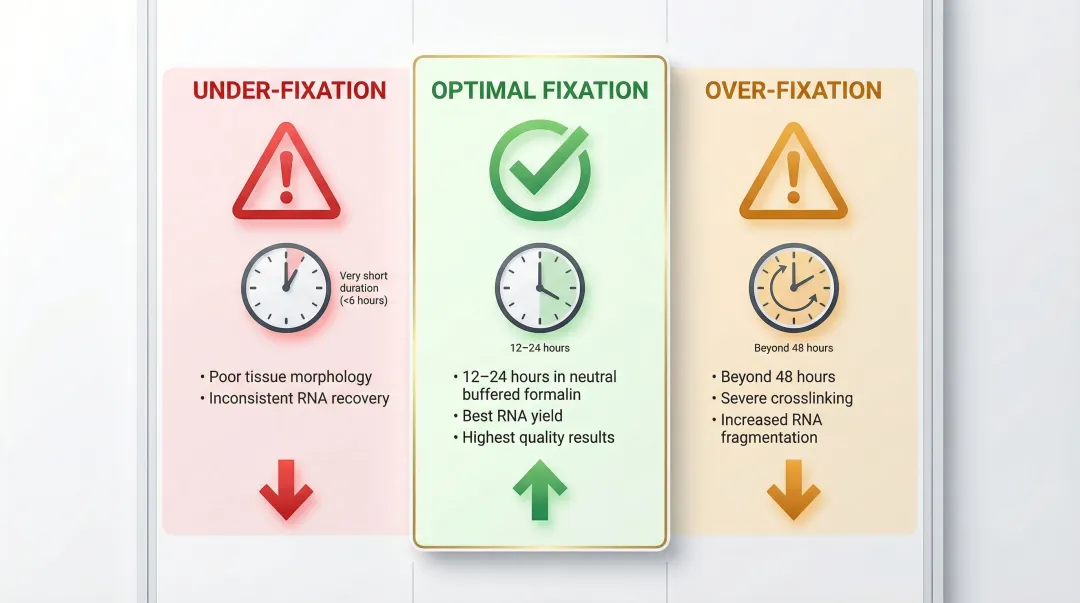
Labs should prioritise samples with documented fixation protocols and avoid blocks with unknown or prolonged fixation histories when possible.
Extracting RNA from FFPE Tissue: A Step-by-Step Protocol Overview
Pre-Extraction Preparation
Section thickness: Cut 6–10 µm sections for optimal RNA yield. Thicker sections (>10 µm) complicate deparaffinisation; thinner sections (<6 µm) reduce total RNA recovery.
Target tissue area: Aim for approximately 18 cm² of tissue (equivalent to roughly 9 slides) to yield sufficient RNA for most sequencing platforms. Adjust based on expected library prep input requirements.
Pathologist-guided macrodissection: Select regions of interest and exclude:
- Necrotic areas (contain degraded RNA and contribute noise)
- Melanin-rich zones (interfere with spectrophotometric quantification)
- Adjacent normal tissue (introduces biological confounders)
Storage before processing: Store cut sections at room temperature in a desiccator to prevent humidity-driven rehydration. Process within 1–2 weeks of cutting for best results.
Deparaffinisation and Rehydration
Remove paraffin completely before proceeding to lysis:
- Xylene or xylene-substitute washes: Two rounds of 5–10 minutes each to dissolve paraffin wax
- Ethanol gradient series: 100%, 95%, 70% ethanol washes to rehydrate tissue
- Air-dry pellets thoroughly before adding lysis buffer
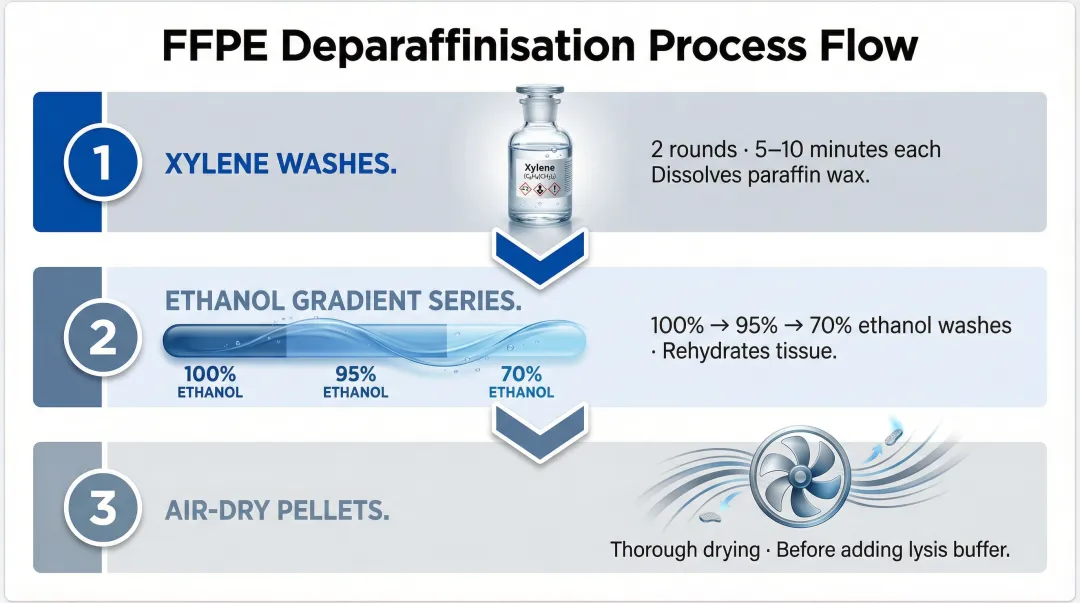

Inadequate deparaffinisation is a common cause of low RNA yield and downstream library failures. Xylene-free deparaffinisation systems — mineral oil-based approaches, for example — have shown improved results for certain tissue types, though xylene-based protocols remain the most widely validated method.
Crosslink Reversal via Proteinase K Digestion
Proteinase K digestion combined with elevated-temperature incubation reverses formalin damage:
- Initial digestion: 56°C incubation to digest protein-RNA adducts
- Heat-mediated reversal: 80°C incubation to reverse methylene bridges chemically
Incubation duration requires optimisation:
- Insufficient incubation leaves crosslinks intact, reducing RNA yield
- Excess incubation increases RNA shearing, lowering fragment length
Most protocols specify 15–60 minutes at 80°C; follow kit-specific recommendations closely.
RNA Purification: Column vs. Bead-Based Methods
Silica membrane spin columns are widely used for FFPE RNA extraction:
- Selective RNA binding to silica in high-salt conditions
- On-column DNase treatment to remove genomic DNA
- Suitable for manual workflows and small-to-moderate sample batches
Magnetic bead-based purification offers advantages for high-throughput settings:
- Automation compatibility with liquid handling systems
- Standardised extraction across sample batches, reducing inter-operator variability
- Reduced hands-on time, particularly valuable for clinical oncology labs processing high volumes
Automated magnetic bead platforms can process 32 FFPE samples in approximately 2.5 hours. For clinical diagnostics labs, that throughput matters: faster, reproducible extractions directly compress the time between sample receipt and actionable sequencing results.
Post-Extraction Yield Expectations and Quantification
Two practical benchmarks guide post-extraction assessment:
- Absorbance yield (NanoDrop): ~400 ng/cm² of tissue — but this figure is inflated by co-purified short nucleotides
- Fluorescence yield (Qubit RNA HS Assay): roughly half the NanoDrop value, reflecting intact RNA more accurately
- Quantification rule: Always use fluorescence-based methods for library prep input calculations; NanoDrop overestimates FFPE RNA concentration by ~2-fold
- Troubleshooting threshold: Absorbance yields below ~246 ng/cm² indicate problems with deparaffinisation or crosslink reversal — revisit those steps before proceeding
With extraction quality confirmed, the next decision is kit selection — where performance differences become especially consequential for degraded FFPE samples.
FFPE RNA Extraction Kit Comparison
Qiagen AllPrep DNA/RNA FFPE Kit
Widely validated across archival sample cohorts up to 20 years old. Enables simultaneous co-extraction of DNA and RNA from the same tissue section, allowing downstream SNP analysis, mutation profiling, or copy number variation studies from residual DNA pellets.
Recommended as the primary choice when both nucleic acid types are required.
- Input: 2–8 tissue sections (10 µm each)
- Protocol time: Approximately 3–4 hours (manual)
- DNase treatment: On-column, included
- Automation compatibility: Compatible with QIAcube
- Downstream compatibility: All major library prep kits
Qiagen RNeasy FFPE Kit
RNA-only extraction optimised for total RNA from FFPE sections. Suitable when DNA is not required and RNA purity is the sole objective. Yields are comparable to AllPrep for RNA; the workflow is simpler because it skips the DNA-binding steps.
- Input: 2–8 tissue sections (10 µm each)
- Protocol time: Approximately 2.5–3 hours (manual)
- DNase treatment: On-column, included
- Automation compatibility: Compatible with QIAcube
- Downstream compatibility: All major library prep kits
TaKaRa NucleoSpin totalRNA FFPE Kit
Suitable for low-input FFPE samples and compatible with the TaKaRa SMARTer library prep workflow, making it an end-to-end solution for labs running low-quantity samples. Published data show average yields of ~127 ng/µL from single 5 µm sections.
- Input: 1–4 tissue sections (5–10 µm each)
- Protocol time: Approximately 2 hours (manual)
- DNase treatment: On-column, included
- Automation compatibility: Limited
- Downstream compatibility: Optimised for TaKaRa SMARTer kits
Zymo Quick-RNA FFPE Kit
Alternative option featuring simplified deparaffinisation with a proprietary reagent system. Reports comparable yields to Qiagen kits with slightly faster protocol times.
- Input: 2–8 tissue sections (10 µm each)
- Protocol time: Approximately 2 hours (manual)
- DNase treatment: On-column, included
- Automation compatibility: Limited
- Downstream compatibility: Broad compatibility
Cambrian FFPE RNA Isolation Kit
Optimised for automated workflows on the Manta platform, this kit uses magnetic bead-based purification with pre-filled cartridges for high-throughput processing. Consistently achieves DV200 >70% and RNA yields of approximately 5,000 ng from just four 10 µm FFPE tissue curls.
- Input: 4 tissue curls (10 µm each)
- Protocol time: Approximately 2.5 hours for 32 samples (automated)
- DNase treatment: Integrated, zero DNA contamination
- Automation compatibility: Native compatibility with Manta (96-well format)
- Downstream compatibility: NGS and PCR applications
Key advantages: Remote monitoring via touchscreen interface, UVC lamp for contamination control, and same-day processing without overnight lysis. Automation also reduces inter-operator variability, which matters when oncology labs are processing dozens of samples per run.
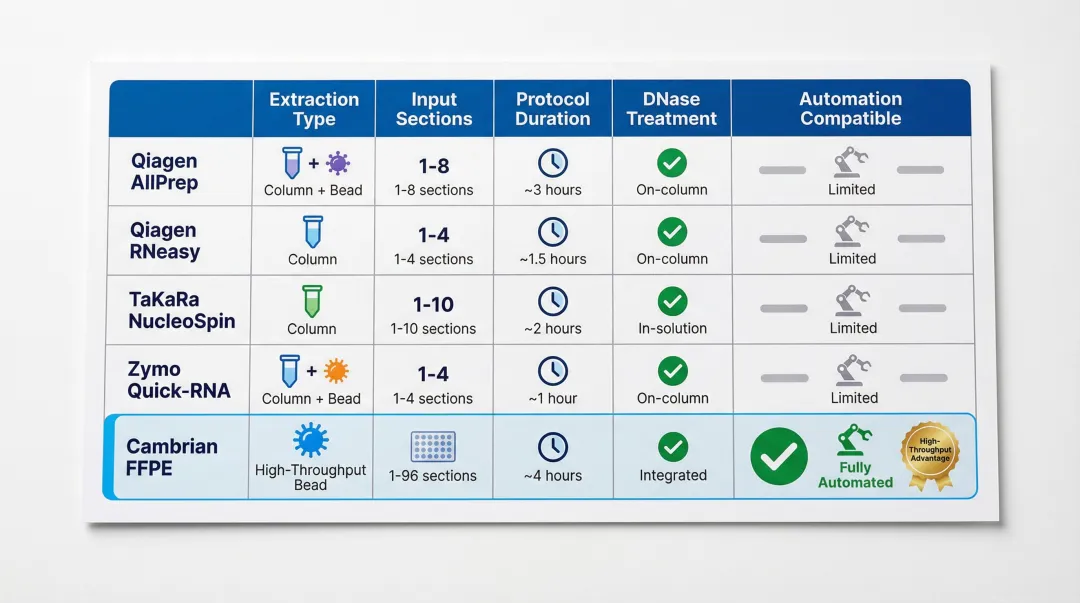
FFPE RNA Extraction Kit Summary Table
| Kit Name | RNA-Only / Dual Extraction | Recommended Input | Reported Yield Range | Protocol Duration | DNase Treatment | Automation Compatible | Downstream Compatibility |
|---|---|---|---|---|---|---|---|
| Qiagen AllPrep DNA/RNA FFPE | Dual (DNA + RNA) | 2–8 sections (10 µm) | 200–600 ng/cm² | 3–4 hours | On-column, included | QIAcube | All major kits |
| Qiagen RNeasy FFPE | RNA-only | 2–8 sections (10 µm) | 200–600 ng/cm² | 2.5–3 hours | On-column, included | QIAcube | All major kits |
| TaKaRa NucleoSpin totalRNA FFPE | RNA-only | 1–4 sections (5–10 µm) | ~127 ng/µL per section | 2 hours | On-column, included | Limited | Optimised for TaKaRa SMARTer |
| Zymo Quick-RNA FFPE | RNA-only | 2–8 sections (10 µm) | 200–500 ng/cm² | 2 hours | On-column, included | Limited | Broad compatibility |
| Cambrian FFPE RNA Isolation | RNA-only | 4 curls (10 µm) | ~5,000 ng per 4 curls | 2.5 hours (32 samples) | Integrated | Manta (96-well) | NGS, PCR |
RIN Thresholds and Why DV200 Is the Right Quality Metric for FFPE RNA
Why RIN Is Inappropriate for FFPE RNA
The RNA Integrity Number (RIN) algorithm was designed to assess intact ribosomal RNA bands (18S and 28S) in fresh or frozen samples. FFPE RNA is universally fragmented, producing RIN values of 1–3 even when samples yield excellent sequencing data.
Do not set a minimum RIN threshold for FFPE samples. Archival FFPE specimens stored up to 20 years routinely report RIN values of 1.2–2.5 while still generating >13,000 protein-coding gene reads. Labs that enforce RIN ≥7 cutoffs for FFPE discard usable material — a costly and avoidable error.
DV200 as the Primary QC Metric
DV200 (percentage of RNA fragments ≥200 nucleotides) measured by Bioanalyzer or TapeStation provides the correct quality assessment for FFPE RNA.
Practical interpretation thresholds:
- DV200 ≥50%: Excellent quality, suitable for all library prep methods
- DV200 30–50%: Workable for most approaches, including standard stranded protocols
- DV200 <30%: High degradation; select degradation-tolerant methods (MACE-seq, TruSeq RNA Exome)
- DV200 <20%: Isn't always a failure point — some samples in the 10–20% range yield acceptable data with optimized protocols
TapeStation Curve Shape Interpretation—Beyond DV200 Alone
The electropherogram curve shape provides additional information not captured by the DV200 number alone:
Samples showing a distinct shoulder or small peak near 100 bp often yield adequate sequencing data even when DV200 appears low. This indicates a population of short but usable fragments.
True "failed" samples show a flat, featureless trace with no peaks—these rarely produce usable libraries regardless of DV200 value.
This nuance helps labs avoid discarding borderline samples that would succeed with appropriate library prep methods.
Fluorescence vs. Absorbance for FFPE RNA Quantification
Once you've assessed RNA quality, accurate quantification determines whether your library prep input is correct. The method you choose here matters.
| Method | Basis | FFPE Accuracy |
|---|---|---|
| NanoDrop (A260) | Absorbance — detects all nucleotides | Overestimates by 2× or more; includes degraded fragments unusable in library prep |
| Qubit RNA HS / Bioanalyzer Pico chip | Fluorescence — selectively binds longer fragments | Accurate measurement of intact, library-prep-relevant RNA |
Always use fluorescence-derived values when calculating library prep input mass. Absorbance-based inputs result in under-loaded libraries, directly reducing sequencing depth and gene detection rates.
Library Preparation and Downstream Data Quality for FFPE RNA-Seq
Choosing the Right Library Prep Kit Based on Input and Goals
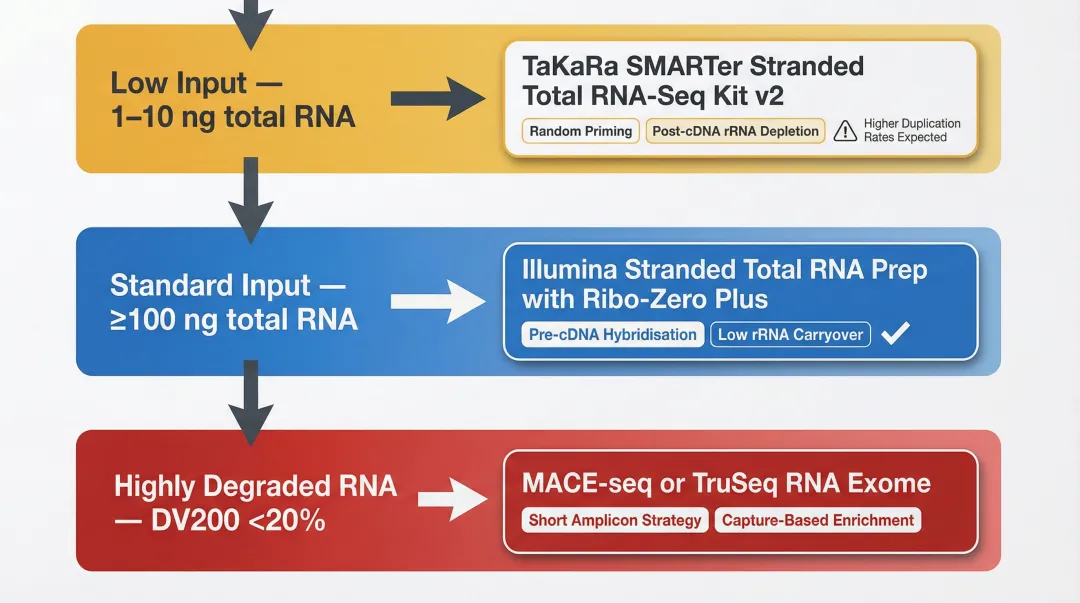
Low RNA input (1–10 ng): TaKaRa SMARTer Stranded Total RNA-Seq Kit v2 uses random priming at the cDNA stage and performs rRNA depletion post-cDNA synthesis, enabling recovery of low-abundance transcripts and fragmented RNA. However, it exhibits higher rRNA carryover (17.45%) and duplication rates (28.48%) per a 2024 FFPE RNA-Seq kit comparison study, requiring deeper sequencing to achieve comparable gene detection.
Standard input (≥100 ng): Illumina Stranded Total RNA Prep with Ribo-Zero Plus delivers superior uniquely mapped read rates and lower rRNA carryover (0.1%). Pre-cDNA hybridisation-based rRNA depletion reduces residual ribosomal contamination more effectively than post-cDNA approaches.
Highly degraded samples (DV200 <20%): 3'-end capture approaches like MACE-seq or capture-based exome kits (Illumina TruSeq RNA Exome) outperform standard whole-transcriptome depletion. These methods rely on shorter amplicon coverage, which tolerates extreme fragmentation better.
Why Poly-A Selection Fails for FFPE and rRNA Depletion Is Mandatory
Poly-A tails are susceptible to formalin-mediated degradation, making mRNA enrichment via oligo-dT selection unreliable. Total RNA sequencing paired with ribosomal RNA depletion (which accounts for >90% of total RNA) is the standard.
Pre-cDNA hybridisation-based Ribo-Zero approaches leave lower residual rRNA but may miss some fragmented transcripts.
Post-cDNA depletion (as in the TaKaRa SMARTer workflow) preserves broader transcript diversity at the cost of slightly higher rRNA carryover.
Key Data Quality Indicators to Monitor Post-Sequencing
Percentage of reads mapping to protein-coding genes:
- TruSeq RNA Exome: ~82%
- MACE-seq: ~88%
- Stranded/Ribo-Zero: ~55% (due to strand-specificity requirements)
GC-content bias is higher in TruSeq libraries compared to Stranded or MACE-seq approaches. Use GC bias tools to flag over-fixed samples as outliers — median absolute deviation (MAD) residuals are more sensitive than PCA for detecting individual technical outliers in FFPE cohorts.
Gene dropout is elevated in low-expression, high-GC genes and is a known artifact of FFPE crosslinking bias. Monitor dropout rates as a routine QC checkpoint.
Sequencing Depth Recommendations
Research suggests 15–20 million mapped reads per sample is sufficient to detect 12,000–16,000 protein-coding genes from FFPE whole-transcriptome profiling.
TaKaRa SMARTer libraries from 5 ng input require higher sequencing depth to achieve gene detection comparable to higher-input Illumina libraries, due to elevated rRNA carryover and duplication rates. Labs should budget for 25–30 million raw reads per TaKaRa sample to compensate.
Cost implications: The sequencing depth penalty for low-input kits adds up quickly. The TaKaRa SMARTer v2 kit carries a lower per-reaction reagent cost (approximately ₹1,100–₹2,300), but the additional sequencing depth required adds roughly ₹5,000–₹8,000 per sample.
Illumina Stranded Total RNA Prep has a higher upfront reagent cost (approximately ₹5,000–₹6,250 per sample), but lower sequencing depth requirements offset this for high-throughput studies. For large cohorts, total project cost often favors the higher-input kit. Note: pricing figures are approximate INR equivalents based on published USD benchmarks.
Frequently Asked Questions
Can you do RNA-seq on formalin-fixed, paraffin-embedded (FFPE) tissue?
Yes, RNA-seq from FFPE tissue is feasible and increasingly common in cancer research. With appropriate extraction protocols, rRNA depletion-based library prep, and quality assessment using DV200 instead of RIN, reliable transcriptomic data can be generated from archival FFPE specimens stored for many years.
What is the difference between FFPE and FF?
FF (fresh frozen) tissue preserves RNA integrity at high quality (typically RIN 7–10) and serves as the gold standard for RNA-seq. FFPE tissue is preserved in formalin and embedded in paraffin for long-term room-temperature storage, resulting in fragmented RNA. FFPE offers far greater sample availability from hospital archives.
What is a good RIN score for RNA-seq?
RIN ≥7 is the standard threshold for fresh/frozen RNA samples. However, this metric is not applicable to FFPE RNA. FFPE samples routinely yield RIN values of 1–3 even when they produce high-quality sequencing data. DV200 is the correct metric to apply to FFPE RNA.
How many million reads are needed for RNA-seq?
For whole-transcriptome FFPE RNA-seq, 15–20 million mapped reads per sample is a common target for detecting 12,000–16,000 protein-coding genes. Read depth requirements are higher for FFPE compared to fresh/frozen samples due to lower RNA integrity and may vary based on library prep method used.
What is a low mapping percentage in RNA-seq?
A uniquely mapped read rate below 50% is generally considered low and warrants investigation. For FFPE RNA-seq, mapping rates vary by library method—stranded Ribo-Zero approaches may show ~55% mapping to protein-coding genes, while capture-based or 3'-end methods can achieve 80–88%. Unusual deviations from expected benchmarks indicate sample quality or library prep issues.
How much does RNA-seq cost per sample?
FFPE RNA-seq costs vary by library prep kit, sequencing depth, and extraction method. A comparative analysis estimates TaKaRa SMARTer v2 at $11–$23 per library prep reaction (approx. ₹900–₹1,900), while Illumina Stranded Total RNA Prep runs $50–$62.50 per sample at academic core facilities. TaKaRa's higher rRNA carryover requires deeper sequencing, which can offset its lower upfront reagent cost.


