Introduction
Cell-free RNA (cfRNA) in plasma has become a clinically relevant non-invasive biomarker source in oncology, prenatal testing, infectious disease monitoring, and liquid biopsy research. Unlike tissue biopsies, plasma sampling offers a minimally invasive window into tissue-specific gene expression changes.
That demand is growing fast. The global cfRNA analysis market is projected to reach $109.16 million by 2032, driven by expanding clinical adoption across molecular diagnostics.
The challenge is that plasma contains exceptionally low cfRNA concentrations—averaging just 7.1 ng/mL in healthy donors and 7.9 ng/mL in cancer patients. That makes extraction method selection critical for downstream accuracy in RT-qPCR, NGS, or small RNA sequencing.
This article covers isolation principles, major extraction methods, and a kit comparison to help labs select the right approach for their specific workflows.
TL;DR
- Cell-free RNA in plasma includes miRNAs, lncRNAs, mRNA fragments, and piRNAs — carried in extracellular vesicles or bound to proteins
- Three main isolation strategies exist: phenol-based (TRIzol), silica spin columns (miRNeasy), and magnetic bead-based automated platforms
- Pre-centrifugation steps, input volume, and elution volume all affect yield and downstream assay performance
- Choose your kit based on target RNA species, intended application (qPCR vs. NGS), and your lab's throughput demands
- Automation reduces manual error and maintains reproducibility in high-volume clinical labs
What Is Cell-Free RNA in Plasma and Why Does It Matter?
Cell-free RNA (cfRNA) in plasma refers to extracellular RNA found outside cells — primarily miRNAs, mRNA fragments, lncRNAs, piRNAs, and tRNA. Despite the name, these molecules are not truly "free": they're shielded from RNase degradation by association with exosomes, microvesicles, apoptotic bodies, lipoproteins, or ribonucleoprotein complexes.
High-throughput sequencing confirms a diverse spectrum of cfRNA biotypes in human plasma, with miRNA fragments typically ranging from 18–22 nt and tRNA fragments showing a dominant size peak at 87 nt.
Protection Mechanisms: More Than Vesicles
While extracellular vesicles (EVs) encapsulate a portion of cfRNA, the majority of circulating miRNAs actually co-fractionate with protein complexes rather than vesicles. Argonaute2 (Ago2) ribonucleoprotein complexes carry up to 90% of circulating miRNAs independent of vesicles, protecting them from protease and RNase activity. Additionally, high-density lipoproteins (HDL) transport endogenous miRNAs in plasma and deliver them to recipient cells.
This molecular stability — maintained across multiple carrier types — is precisely what makes cfRNA detectable and clinically useful from a simple blood draw.
Biomarker Significance
Circulating cfRNA profiles reflect tissue-specific gene expression changes and have been linked to:
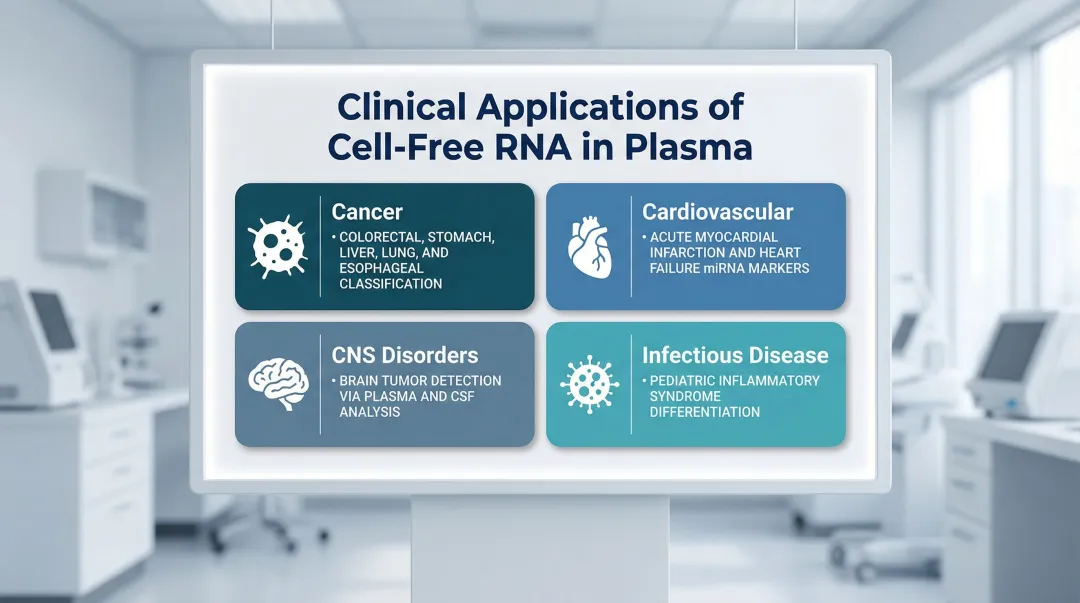
- Cancer: Plasma cfRNA classifies multiple cancer types — colorectal, stomach, liver, lung, and esophageal — and distinguishes malignant from pre-malignant conditions
- Cardiovascular disease: Circulating miRNAs serve as biomarkers for acute myocardial infarction and heart failure
- CNS disorders: Plasma and CSF cfRNA signatures show strong performance in detecting brain tumors
- Infectious disease: cfRNA profiles differentiate pediatric inflammatory syndromes, including Kawasaki disease and MIS-C
Plasma vs. Serum: Why Plasma Is Preferred
Plasma (collected with anticoagulants like EDTA) is strictly preferred over serum for cfRNA studies. Serum preparation involves clot formation, which induces platelet activation and degranulation, releasing intracellular RNA ex vivo that skews the circulating RNA profile. This platelet-derived contamination confounds biomarker studies — making EDTA plasma the recommended matrix for reproducible cfRNA profiling.
Methods for Isolating Cell-Free RNA from Plasma
Organic/Phenol-Based Extraction (TRIzol/TRIzol LS)
Phenol-guanidinium lysis releases RNA through phase separation and precipitation. TRIzol LS is adapted for liquid samples, but standard TRIzol protocols are designed for long RNA and may underperform for small non-coding RNAs (below 60 nt).
TRIzol Frozen Precipitation (TFP) improves recovery of very short sncRNAs (30–60 nt) that column-based kits routinely miss. The protocol uses:
- Phenol-based lysis at room temperature
- Precipitation with isopropyl alcohol, glycogen (2 µL), and 3 M sodium acetate (50 µL)
- Freezing at −80°C for ~25 minutes
TFP yields CT values 2.01–9.17 cycles lower for 30–60 nt sncRNAs compared to standard column kits.
Drawbacks: Labor-intensive, operator-dependent, and prone to organic solvent carryover that inhibits downstream RT-qPCR.
Silica Membrane Spin-Column Methods
RNA binds to silica membranes under chaotropic salt conditions, is washed to remove contaminants, and eluted in a small volume. These methods (miRNeasy, Norgen, miRCURY) are widely used for miRNA isolation.
Size Limitations: Standard silica columns fail to efficiently capture RNA shorter than 200 nt without optimized chemistry. Specialized kits capture RNA >18 nucleotides, but columns routinely lose ultra-short RNAs due to size-dependent binding efficiency and wash-related loss.
Vesicle Enrichment Pre-Isolation Steps
Before RNA extraction, labs can enrich extracellular vesicles using:
- Precipitation (ExoQuick/PEG-NaCl): Simple, fast, high EV and RNA yield; co-precipitates non-EV proteins (albumin) and lipoproteins
- Membrane affinity columns (ExoRNeasy): Fast workflow (~1 hour), concentrates EVs from up to 4 mL plasma; excludes non-vesicular RNP-bound circulating RNAs (e.g., Ago2-bound miRNAs)
- Size-exclusion chromatography (SEC): High purity, preserves vesicle integrity, removes soluble proteins; requires sample concentration
- Differential ultracentrifugation: No additional chemicals, isolates from large volumes; low RNA yield, co-isolates non-exosomal impurities
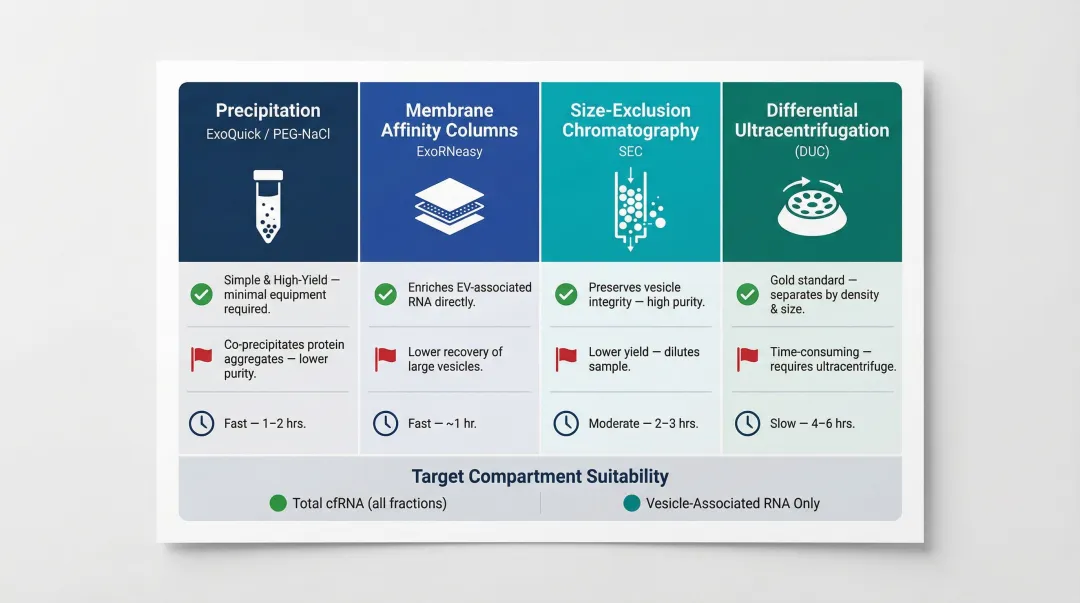
Choose the method based on your target compartment: precipitation and ultracentrifugation capture total cfRNA from whole plasma, while membrane affinity columns restrict analysis to vesicle-associated RNA.
Magnetic Bead-Based Extraction
Once EVs are enriched (or when working from whole plasma), magnetic bead-based extraction offers a direct path to automation. Paramagnetic beads coated with specific ligands or chaotropic chemistry capture RNA from lysed plasma, which is then washed and eluted in a defined volume. The format integrates readily with liquid handling platforms, supporting consistent, hands-free processing across batch sizes — from a handful of samples to full 96-well runs in a clinical setting.
Scaling Plasma Input Volumes for NGS
Standard kits use 200 µL plasma, which yields insufficient RNA for library preparation. Studies show that scaling to 9 mL plasma using vacuum manifold systems (e.g., QIAvac24 Plus with miRNeasy) yielded 77.4 ± 24.9 ng total RNA—successfully generating the ~35 ng required for small RNA-Seq library preparation. Labs must balance increased volume against column clogging, centrifugation time, and sample availability.
Top RNA Extraction Kits for Plasma: A Comparison
The kits below are selected based on published validation data, use in peer-reviewed protocols, downstream application flexibility, and suitability for clinical and research settings. Performance varies by target RNA species, input volume, and downstream application.
Qiagen miRNeasy Serum/Plasma Kit (Advanced)
Industry-standard silica column kit designed specifically for miRNA and total cell-free RNA isolation from 200 µL serum or plasma. Widely validated in published research and cited as the reference method in multiple comparative studies.
Broad RNA size coverage (18–200 nt optimally), compatibility with downstream qRT-PCR and small RNA-Seq, and a high-throughput format using the QIAvac24 Plus vacuum manifold make it scalable to 9 mL plasma volumes for NGS applications.
| Feature | Details |
|---|---|
| Method | Silica membrane spin column with QIAzol phenol lysis |
| Input Volume | 200 µL plasma (scalable to 9 mL with vacuum manifold) |
| Best For | miRNA isolation, qRT-PCR, small RNA sequencing |
Norgen Plasma/Serum Circulating and Exosomal RNA Purification Kit
Column-based kit using proprietary resin chemistry (not silica) designed to co-purify both vesicle-associated and free circulating RNA from 200–500 µL plasma without requiring a separate vesicle enrichment step.
The simplified workflow requires no chloroform or phenol, handles both miRNA and larger RNA species, and includes an optional Amicon filtration step for concentration. Noted in comparative data to show variable yields depending on sample type and RNA target.
| Feature | Details |
|---|---|
| Method | Proprietary resin column (no organic solvents) |
| Input Volume | 200–500 µL plasma |
| Best For | Simplified cfRNA isolation; labs avoiding organic reagents |
Qiagen miRCURY RNA Serum & Plasma Kit (Formerly Exiqon)
Spin-column kit using a unique proprietary buffer system with isopropanol precipitation that efficiently captures small RNAs including miRNAs from 200–500 µL serum/plasma. Originally developed by Exiqon and widely used for circulating miRNA profiling applications.
Strong performer for miRNA recovery with streamlined protocol that avoids chloroform; performs well in comparative studies for consistent miRNA yield and purity, making it a reliable option for clinical biomarker assay development.
| Feature | Details |
|---|---|
| Method | Spin column with proprietary lysis and isopropanol precipitation |
| Input Volume | 200 µL plasma |
| Best For | Circulating miRNA biomarker studies; standardized clinical research |
Promega Maxwell RSC miRNA Plasma and Serum Kit
Automated cartridge-based kit designed for the Maxwell RSC or Maxwell RSC 48 instruments, processing up to 16 or 48 samples in approximately 70 minutes using pre-dispensed reagent cartridges.
Full automation from lysis to elution reduces technician hands-on time, minimises inter-run variability, and delivers concentrated RNA eluate in low volume. Well-suited for high-throughput clinical labs requiring reproducibility and traceability.
| Feature | Details |
|---|---|
| Method | Automated paramagnetic particle-based extraction (cartridge system) |
| Input Volume | 200 µL plasma/serum |
| Best For | High-throughput clinical labs; standardised qRT-PCR workflows |
Cambrian Bioworks Manta Automated Extraction System
Compact, automated nucleic acid extraction platform using magnetic bead-based chemistry for DNA, RNA, and cell-free nucleic acid purification—designed for clinical and research labs that need reliable, walkaway extraction without the footprint of large liquid handlers.
The Manta runs extractions in approximately 30 minutes, operates without batching pressure, and fits inside a biosafety hood—practical advantages for oncology, liquid biopsy, and molecular diagnostics labs where operator variability and throughput flexibility both matter. ISO 13485-certified and CE-IVD marked, it is trusted by 67+ labs globally.
| Feature | Details |
|---|---|
| Method | Magnetic bead-based automated extraction with pre-filled cartridges |
| Input Volume | Compatible with diverse sample types including plasma |
| Best For | Clinical and research labs needing automated, reproducible cfRNA/cfDNA extraction with compact instrument footprint |

How to Choose the Right RNA Extraction Method for Your Lab
Target RNA Species Is the First Decision Point
If the goal is miRNA profiling, column-based kits (miRNeasy, miRCURY) and automated magnetic bead platforms are well-validated. For recovering very short sncRNAs (30–60 nt such as pfeRNAs or piRNAs), standard column methods may miss these due to size-dependent binding efficiency—consider optimisation (such as glycogen precipitation or modified elution) or organic-based protocols. For whole transcriptome or mRNA fragment profiling, higher plasma input volume and NGS-optimised workflows are required.
Downstream Application Drives Method Selection
For RT-qPCR, most commercial kits provide sufficient quality. For NGS or small RNA-Seq, the extremely low RNA yield from standard 200 µL input is typically insufficient—researchers should consider scaling plasma input, using vacuum manifold systems, or fluorometric quantification (Qubit, for example) rather than NanoDrop, which often reads below detection for cfRNA concentrations.
Fluorometric assays provide the lowest coefficient of variation (5.47%) for low-abundance plasma RNA, whereas spectrophotometric methods overestimate concentrations by detecting phenol, silica carryover, and protein contaminants.
Throughput and Automation Requirements
Manual protocols introduce inter-operator variability and are not scalable in high-volume diagnostic settings. Labs processing more than 20 samples per day or requiring CAP/CLIA-level reproducibility should prioritise automated platforms with pre-filled reagent cartridges and onboard quality controls. Automated systems also reduce contamination risk through enclosed processing.
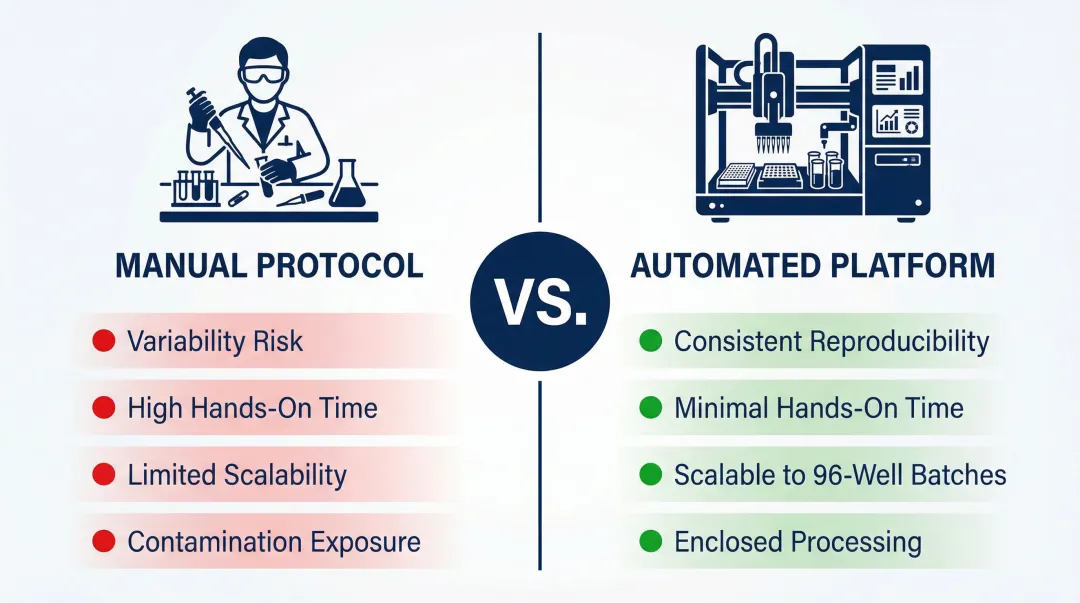
Choosing the right automation platform also depends on getting pre-analytical steps right — even the best extraction method fails if sample handling is inconsistent.
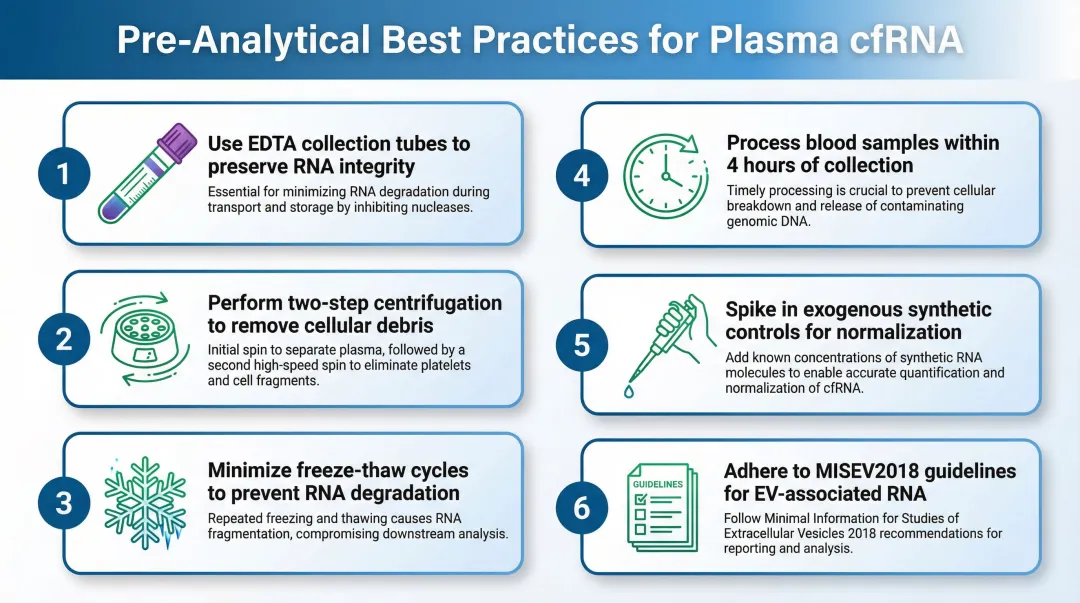
Pre-Analytical Variables Are Commonly Overlooked
Each variable below independently affects RNA yield and downstream data quality:
- Use EDTA plasma, not heparin — heparin strongly inhibits reverse transcriptase and PCR enzymes, compromising downstream analysis.
- Apply two-step centrifugation — spin first at ~1,600×g, then at ~16,000×g to clear cells and debris before extraction begins.
- Minimise freeze-thaw cycles — repetitive freeze-thaw significantly reduces plasma miRNA concentrations and distorts RNA profiles. Store at −80°C and process in a single thaw.
- Process whole blood promptly — plasma should be separated within 4 hours of collection to prevent leukocyte and erythrocyte lysis from contaminating the fraction.
- Spike in exogenous controls — synthetic miRNAs (such as C. elegans cel-miR-39) added to lysis buffer at known concentrations monitor extraction efficiency and normalise RT-qPCR data.
- Follow ISEV reporting standards — the MISEV2018 guidelines mandate reporting of biofluid source volumes, EV separation specificity, and both positive and negative markers for co-isolated non-EV components.
Conclusion
Cell-free RNA isolation from plasma remains one of the most technically demanding steps in liquid biopsy workflows. Method choice must align with target RNA type, downstream application, throughput demands, and lab infrastructure. No single kit is universally optimal across all use cases.
Validate any kit internally with spike-in controls and appropriate quantification methods before committing to a clinical workflow. Consider whether automation is a priority for consistency at scale.
For labs where automation is the answer, Cambrian Bioworks offers compact magnetic bead-based extraction platforms designed for RNA and cell-free nucleic acid workflows—trusted by 67+ labs across clinical and research settings, with ISO 13485 and CE-IVD certification. Their team can help evaluate compatibility with your plasma RNA extraction setup at connect@cambrianbioworks.com.
Frequently Asked Questions
Can you extract RNA from plasma?
Yes. Plasma contains cell-free extracellular RNA (cfRNA) at very low concentrations, protected within extracellular vesicles or bound to proteins. Specialised kits or optimised protocols are required to achieve usable yields for downstream molecular applications.
What does EDTA do to RNA?
EDTA chelates divalent cations (Mg²⁺, Ca²⁺), which inhibits RNase activity and helps preserve RNA integrity in blood collection tubes. EDTA plasma is the preferred anticoagulant for cfRNA studies, whereas heparin should be avoided as it strongly inhibits reverse transcriptase and PCR enzymes.
Can I freeze cells before RNA extraction?
For plasma cfRNA specifically, freezing processed (post-centrifugation, cell-free) plasma at −80°C is acceptable for short-term storage. Repeated freeze-thaw cycles degrade cfRNA, so each aliquot should be used in a single thaw.
What is the difference between plasma and serum for RNA extraction?
Plasma is obtained before clotting (with anticoagulant) and retains platelets, while serum is collected after clotting and platelet activation. Platelet activation during clotting releases intracellular RNA that alters the circulating RNA profile. For cfRNA biomarker studies, plasma is generally preferred.
How do I prevent RNA degradation during plasma sample handling?
Process blood within 2–4 hours of collection, perform two-step centrifugation to remove cells and debris, avoid haemolysis (which releases cell-derived RNA), store plasma aliquots at −80°C, and minimise freeze-thaw cycles. RNase inhibitors are not added to plasma prior to extraction.
Which extraction method is best for miRNA isolation from plasma?
Silica column-based kits such as the Qiagen miRNeasy Serum/Plasma Kit and Qiagen miRCURY RNA Serum & Plasma Kit are among the most widely validated for circulating miRNA isolation. Automated magnetic bead platforms offer equivalent or better reproducibility at scale, with the final choice driven by throughput needs, downstream application, and whether a vesicle enrichment pre-step is required.