Introduction
A lung cancer patient receives a liquid biopsy result: EGFR T790M mutation negative. Treatment continues, but the tumor progresses. A follow-up tissue biopsy reveals the mutation was present all along—at 0.3% variant allele fraction (VAF). The false negative didn't occur because the mutation wasn't there. It happened because the extraction protocol lost most of the circulating tumor DNA (ctDNA) before the assay could detect it.
This scenario repeats across oncology labs worldwide. Liquid biopsy sensitivity depends on a chain of steps: blood draw → plasma separation → cfDNA extraction → library prep → sequencing or PCR. Among these, extraction quality is the earliest variable a laboratory can directly control — and the one most frequently under-optimized.
In early-stage cancers, ctDNA can represent as little as 0.01% of total cfDNA. Even modest extraction losses push these samples below detection limits, turning true positives into false negatives.
Understanding where those losses occur — and how to prevent them — is what separates a reliable liquid biopsy program from one that misses the mutations that matter most. This article examines three critical dimensions of cfDNA extraction quality: yield, fragment integrity, and purity, explaining how each affects downstream assay performance and what validated, reproducible workflows look like in practice.
TLDR
- cfDNA concentrations are extremely low (1–10 ng/mL healthy; ~12.6 ng/mL in cancer patients), so small extraction losses push template below detection limits
- gDNA contamination from lysed white blood cells dilutes ctDNA variant allele fractions, masking true tumour mutations
- Fragment integrity matters: cfDNA exists as ~166 bp nucleosomal fragments; degradation reduces library prep efficiency and read quality
- Pre-analytical errors (delayed plasma separation, wrong tube type, freeze-thaw cycles) compromise cfDNA quality before extraction even begins
- Automation and validated extraction kits reduce inter-run variability and are increasingly required for clinical-grade workflows
Why cfDNA Extraction Quality Is the Bottleneck in Liquid Biopsy Sensitivity
cfDNA is uniquely challenging to extract. It circulates at picomolar concentrations, and ctDNA specifically constitutes only 0.01–1% of total cfDNA in cancer patients. This creates an acute signal-to-noise problem: any process-introduced loss disproportionately impacts assay sensitivity.
The Molecule Math of Low VAFs
Consider the numbers. A standard 10 mL blood draw yields approximately 5 mL of plasma, containing roughly 50 ng of total cfDNA—about 15,000 haploid genome equivalents. At a 0.1% VAF (typical for localized malignancy or minimal residual disease), this translates to just 15 mutant DNA molecules. If extraction efficiency drops by 50%, you lose half of those molecules, effectively doubling the limit of detection (LOD). The assay no longer sees the mutation.
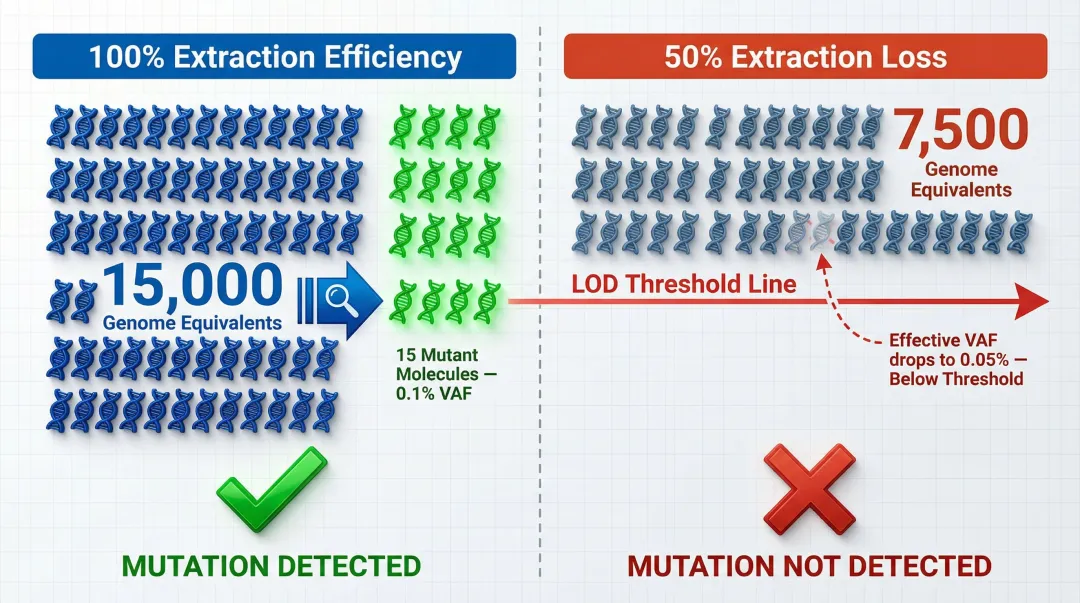
The clinical consequence is a false negative indistinguishable from a true one — meaning a patient may continue an ineffective therapy or have early relapse go undetected.
Why Tissue DNA Extraction Doesn't Face This Problem
In tissue biopsy, DNA is abundant—micrograms per sample. A 20–30% extraction loss is inconsequential; plenty of template remains. In plasma cfDNA workflows, where input is measured in nanograms, the same relative loss renders samples unanalyzable.
Quantitative loss is only part of the problem. The chemistry of extraction also introduces a fragment-size bias that selectively disadvantages the ctDNA signal.
Fragment Size Bias: Not All Extraction Methods Recover cfDNA Equally
Every extraction workflow passes samples through the same core steps — lysis, separation, binding, washing, and elution — and each is a potential loss point for short cfDNA fragments.
Short fragments bind less efficiently to silica columns than high-molecular-weight gDNA. Conventional silica-based kits show limited recovery of fragments below 150 bp—recovering only 18% of 150-nucleotide fragments and virtually none (0.2%) of 40-nucleotide fragments in spiked urine studies. Conversely, magnetic bead-based methods generally recover a higher fraction of short cfDNA fragments, making them better suited than column-based methods for liquid biopsy workflows.
Because ctDNA is characteristically shorter than background cfDNA from healthy cells, gDNA-optimized extraction methods systematically under-recover the very molecules the assay needs to detect.
The Three Pillars of cfDNA Quality: Yield, Fragment Integrity, and Purity
Yield
cfDNA yield depends on plasma volume, concentration, and elution conditions. Expected concentration ranges vary by tumor type and burden:
- Healthy individuals: 1–10 ng/mL
- Cancer patients (stage I-III): ~12.6 ng/mL average
- Lung cancers: ~5.23 ng/mL; liver cancers: ~46.0 ng/mL
Insufficient yield creates template insufficiency downstream. For digital droplet PCR (ddPCR), fewer droplets contain mutant molecules, reducing quantification precision. For NGS, low input reduces library complexity — unique template molecules are at higher risk of being lost during PCR amplification.
Extraction and elution volumes directly affect final yield concentration. Labs typically process 1–4 mL plasma and elute in 50–200 µL. Smaller elution volumes increase DNA concentration but require highly efficient binding and release steps to avoid losses during elution.
Fragment Integrity
cfDNA's characteristic nucleosomal fragment size is approximately 167 bp, with di-nucleosomal peaks around 332 bp. This pattern functions as a diagnostic signature and a direct quality indicator.
Degraded cfDNA with a smeared profile below 100 bp indicates nuclease activity from delayed processing or suboptimal extraction. Fragment integrity affects specific assay types differently:
- NGS library prep requires fragments with intact ends for adapter ligation; degraded cfDNA produces short, unusable library molecules
- qPCR amplicons spanning 100–200 bp need a minimum number of intact template molecules
- Methylation-based assays depend on fragment integrity for accurate bisulfite conversion without bias
Purity
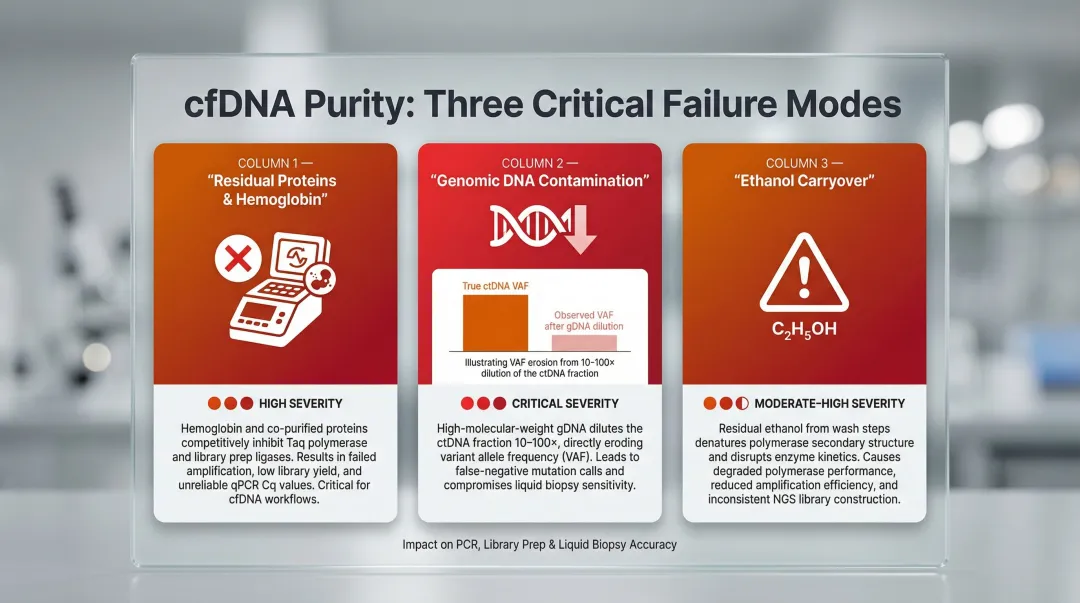
Three main purity concerns affect cfDNA extracts:
1. Residual proteins/hemoglobin that inhibit PCR and library prep enzymes
2. Co-eluted genomic DNA from white blood cells that dilutes the ctDNA fraction
3. Residual wash buffer carryover (ethanol) that degrades polymerase performance
gDNA contamination is the most critical purity issue. Plasma samples held for more than 4–6 hours at room temperature without cell stabilisation risk substantial white blood cell lysis. The released gDNA can increase total DNA input 10–100 fold, proportionally diluting the ctDNA fraction's representation in any downstream assay.

How Extraction Failures Translate to False Negatives
Scenario: EGFR T790M at 0.3% VAF
A lung cancer patient has an EGFR T790M resistance mutation at 0.3% VAF. The mutation is present, but extraction recovers only 30% of cfDNA input due to column binding inefficiency. The effective VAF presented to the ddPCR assay drops to ~0.09%—below the assay's 0.1% LOD. The result is reported as negative, and the patient continues on an ineffective therapy.
The gDNA Dilution Effect
If extraction produces 20 ng cfDNA at 0.3% VAF but also co-elutes 50 ng gDNA (wild-type), total DNA input is now 70 ng. The EGFR T790M molecules are diluted to approximately 0.09% VAF—again, below the assay's LOD.
Consequences for NGS
These upstream extraction failures compound downstream in sequencing workflows. Degraded cfDNA produces short adapter-ligated fragments that are either too small to cluster on flow cells or fail alignment quality filters as extremely short reads. The result: reduced library complexity and specific genomic regions dropping below minimum coverage depth for variant calling.
Inter-Laboratory Reproducibility Problems
When different labs using different extraction methods analyze split samples from the same patient, variant detection concordance can be poor—not because the assays differ, but because extraction-driven VAF differences push borderline-positive samples to different sides of the LOD.
In a pooled analysis of the AURA extension and AURA2 studies for advanced NSCLC, plasma testing detected the EGFR T790M mutation in only 61% of patients confirmed positive by tissue testing—meaning nearly 4 in 10 patients with a confirmed resistance mutation received a false negative result from plasma, with direct consequences for treatment decisions.
Pre-Analytical Variables That Compromise cfDNA Before Extraction Begins
Blood Tube Selection
- EDTA tubes are standard but allow cell lysis within 4–6 hours at room temperature, releasing gDNA
- Cell-free DNA preservation tubes (Streck Cell-Free DNA BCT) stabilise cell membranes for up to 14 days at 6°C to 37°C, preventing nucleated blood cell lysis and gDNA release
- Trade-off: Preservation tubes require assay-specific validation and cost more, but significantly reduce pre-analytical variability
Time-to-Centrifugation
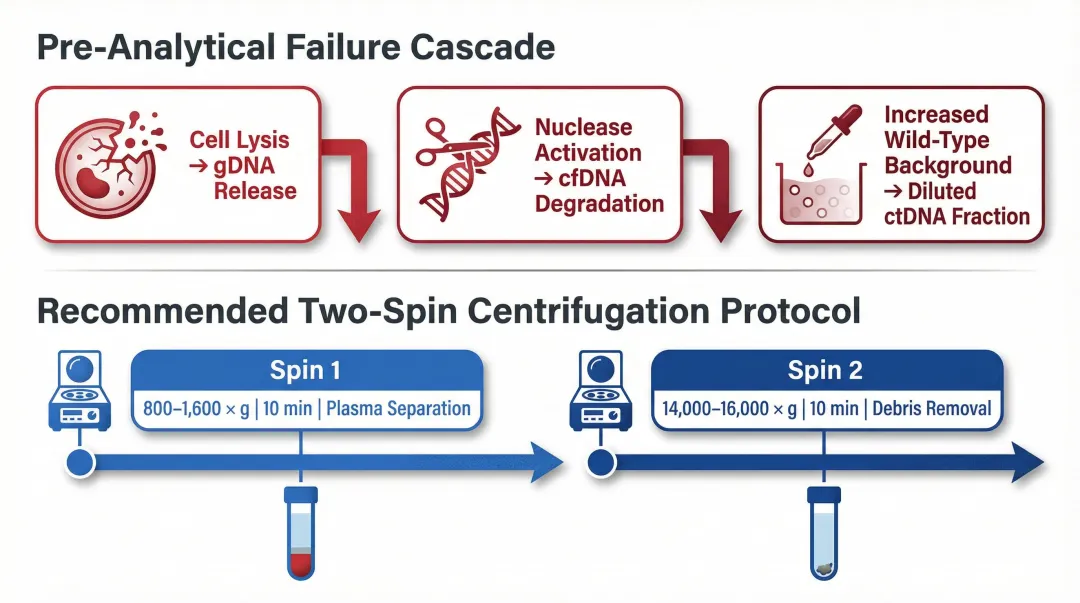
The most impactful pre-extraction variable. Delayed processing triggers a cascade:
- Cell lysis → gDNA release
- Nuclease activation → cfDNA degradation
- Increased background wild-type DNA → diluted ctDNA fraction
Guidelines recommend plasma separation within 4–6 hours using a two-spin protocol:
- First spin: 800–1,600 × g for 10 minutes (separates plasma from cells)
- Second spin: 14,000–16,000 × g for 10 minutes (removes remnant cells and debris)
Freeze-Thaw Cycling
Once plasma is separated and stored, freeze-thaw cycling becomes the next point of failure. Each cycle introduces mechanical shearing of cfDNA fragments and can activate latent nucleases — progressively degrading the very material downstream assays depend on.
To avoid this:
- Aliquot plasma into single-use volumes immediately after centrifugation
- Store aliquots at -80°C and thaw only what is needed per run
Manual vs. Automated Extraction: What the Data Says
Inter-Operator Variability in Manual Extraction
Manual column-based extraction introduces variability through:
- Pipetting technique differences
- Inconsistent spin speeds or durations
- Variable wash step timing
- Elution temperature fluctuations
These operator-dependent variables are difficult to standardise across shifts, operators, or lab sites. For low-input samples where small absolute losses have large relative impacts, this variability directly degrades assay sensitivity.
One study reported a median coefficient of variation (CV) of 11.8% for manual QIAamp kits, 16.0% for automated MagNA Pure systems, and up to 34.6% for other magnetic bead kits.
Automated Liquid Handling Platforms
Closed-system extraction using pre-validated protocols, fixed volumes, and controlled timing removes operator-dependent variables. This is critical for low-input samples.
When evaluating automated platforms for cfDNA work, the key criteria are fragment-size selectivity, gDNA exclusion, throughput flexibility, and validated clinical-grade chemistry. Cambrian Bioworks' Manta was designed around these requirements.
Regulatory & quality credentials:
- CE-IVD marked for clinical DNA extraction under EU Regulation 2017/746 (IVDR)
- ISO 13485-certified design and manufacturing
Extraction performance:
- Magnetic bead chemistry captures short cfDNA fragments (150–200 bp) more consistently than silica columns
- Minimal gDNA co-elution, preserving ctDNA fraction integrity
- Integrated fluorometer for automated post-extraction quantification
Workflow & throughput:
- ~55-minute total run time from plasma to purified cfDNA
- 1–32 samples per run, eliminating batching pressure
- Open-system compatibility for NGS, ddPCR, and qPCR downstream workflows
In practice, the magnetic bead chemistry's preferential capture of short fragments — combined with low gDNA carry-over — directly addresses the two variables most likely to erode assay sensitivity in ctDNA detection.

Best Practices to Standardise cfDNA Extraction Quality in Clinical Labs
Extraction quality failures rarely announce themselves — they surface as unexplained sensitivity drops or failed runs. These three practices build in the controls needed to catch problems before they reach the assay.
Implement a Quality Control Gate Post-Extraction
Assess every cfDNA extract before proceeding to downstream assays:
Quantification:
- Use fluorometric methods (dsDNA-specific assays like Qubit) rather than qPCR-based quantification, which can over-estimate yield when gDNA is present
Fragment size distribution:
- Assess using Bioanalyser or TapeStation to confirm the ~166 bp nucleosomal peak and detect high-molecular-weight gDNA contamination
Define a Validated cfDNA Extraction SOP
A validated SOP should specify each variable that affects yield and purity:
- Plasma volume input — typically 1–4 mL, defined per assay requirement
- Tube type and processing window — validated combination with a maximum hold-time limit
- Extraction kit or platform — documented recovery rate, gDNA contamination level, and CV
- Acceptance criteria — defined yield and purity thresholds that must be met before the assay proceeds
Validate with Contrived Samples
Extraction method validation should be performed on contrived samples with known ctDNA VAFs (for example, Seraseq or Horizon reference standards) to establish method-specific limits of detection, rather than relying on kit manufacturer specifications alone. The BloodPAC consortium and AMP/CAP guidelines are explicit: LOD must be determined from your actual extraction workflow, not inferred from kit datasheets. Manufacturer specifications rarely account for site-specific variables like plasma quality or processing delays.
Frequently Asked Questions
What is a liquid biopsy?
A liquid biopsy is a minimally invasive blood test that detects cancer-related biomarkers—including cfDNA, ctDNA, and circulating tumour cells—circulating in the bloodstream, offering an alternative or complement to tissue biopsy for cancer diagnosis and monitoring.
What biomarkers (ctDNA, cfDNA, CTCs) do liquid biopsies analyse?
cfDNA is all cell-free DNA in circulation; ctDNA is the cancer-derived fraction carrying tumour-specific mutations; CTCs are intact cancer cells shed from tumours. cfDNA/ctDNA is currently the most clinically deployed biomarker.
What type of sample is needed for a liquid biopsy?
Most liquid biopsies use a peripheral blood draw (typically 10–20 mL), from which plasma is separated. The type of collection tube and processing speed both significantly affect the quality of cfDNA recovered for analysis.
What does a negative liquid biopsy result mean?
A negative result does not definitively exclude cancer. It may reflect true absence of detectable disease, a tumour shedding very little cfDNA, or a false negative from assay sensitivity limitations or poor extraction quality. Tissue biopsy confirmation is standard practice when liquid biopsy is negative.
What is the difference between a tissue biopsy and a liquid biopsy?
Tissue biopsy removes cells directly from a tumour for histologic and molecular analysis — the gold standard for diagnosis. Liquid biopsy non-invasively samples tumour-derived DNA from blood and cannot replace tissue biopsy for initial diagnosis, but offers clear advantages for serial monitoring.
How long do liquid biopsy results take?
Turnaround time depends on assay type: rapid ddPCR assays typically return results in 4–6 days, while comprehensive NGS panels report in around 7 days. Extraction and QC add several hours upstream. Automated platforms like Manta cut pre-analytical processing time to under an hour.


