Introduction
Every downstream assay — PCR, qPCR, NGS — is only as reliable as the nucleic acid input it receives. When extraction quality is compromised, results follow.
Magnetic bead-based separation addresses this directly. It uses magnetically responsive particles coated with nucleic acid-binding surfaces to selectively capture, purify, and elute DNA or RNA from complex biological samples — with a consistency that manual and column-based methods struggle to match.
This article is written for clinical and research labs, sequencing facilities, and molecular diagnostics teams who need more than a surface-level overview. It covers the mechanism, the step-by-step workflow, the variables that affect output quality, and where magnetic bead technology is — and is not — the right fit.
TL;DR
- Magnetic beads bind nucleic acids via surface chemistry, immobilize them under a magnetic field, and release purified material after contaminants are washed away
- The process is fast (typically under 30 minutes), automation-compatible, and scalable for high-throughput clinical and research settings
- Yield and purity depend on bead coating chemistry, lysis conditions, wash stringency, and elution parameters
- Compatible with DNA, RNA, and cell-free DNA extraction from blood, plasma, tissue, swabs, and FFPE samples
- Protocol optimisation is required — this is not a universal replacement for all extraction methods
What Is Magnetic Bead-Based Separation for Nucleic Acid Extraction?
Magnetic beads are typically iron oxide cores coated with a nucleic acid-binding surface—most commonly silica. When introduced into a lysed biological sample, they selectively capture DNA or RNA through surface chemistry interactions, then are immobilized by an external magnetic field so contaminants can be washed away cleanly.
The result is purified, inhibitor-free nucleic acid in a defined elution volume, ready for use in PCR, qPCR, next-generation sequencing, or other downstream molecular assays.
How it differs from alternatives:
- Spin column methods rely on centrifugal force and fixed membrane geometry, limiting scalability and automation
- Phenol-chloroform extraction uses chemical phase separation — manual, hazardous, and difficult to standardize
- Salting-out precipitation avoids harsh chemicals but requires multiple high-speed centrifugation steps and is hard to automate
Magnetic bead-based separation eliminates these bottlenecks, making it the preferred format for labs that need consistent, scalable results — whether processing 8 samples or 96.

Why Clinical and Research Labs Use Magnetic Bead-Based Extraction
Automation Compatibility Drives Adoption
Because bead binding and release are controlled entirely by applying or removing a magnetic field—with no centrifuge, column, or filtration step—the workflow integrates directly into liquid handling instruments. This enables unattended batch processing and frees lab staff for higher-value tasks. Automated platforms can process up to 96 samples in under 45 minutes, saving hours of manual labor per day.
Consistency and Reproducibility for IVD-Grade Applications
For IVD-grade applications, batch-to-batch variation in extraction must be minimized. Magnetic bead workflows are highly standardizable compared to membrane-based methods, which are sensitive to centrifuge speed variability and membrane lot differences. Clinical validation studies show CV rates of <4% for Ct values and <2.1% variance in cfDNA extraction yields.
Sample Type Versatility
A single bead-based platform can handle:
- Whole blood, plasma, serum
- FFPE tissue sections
- Nasopharyngeal swabs
- Saliva and urine
- Cell cultures
This versatility eliminates the need for multiple extraction platforms when labs run multiple test menus.
Turnaround Time and Purity Benefits
Magnetic bead-based extraction reduces hands-on time and total protocol duration versus manual spin column workflows, particularly in high-volume environments.
Bead-based wash steps are also highly effective at clearing PCR inhibitors such as heme, lipids, and salts. This matters because incomplete removal of chaotropic salts or ethanol can delay qPCR Ct values or completely inhibit amplification, so optimized wash protocols directly determine whether downstream assays succeed or fail.
How Magnetic Bead-Based Nucleic Acid Extraction Works
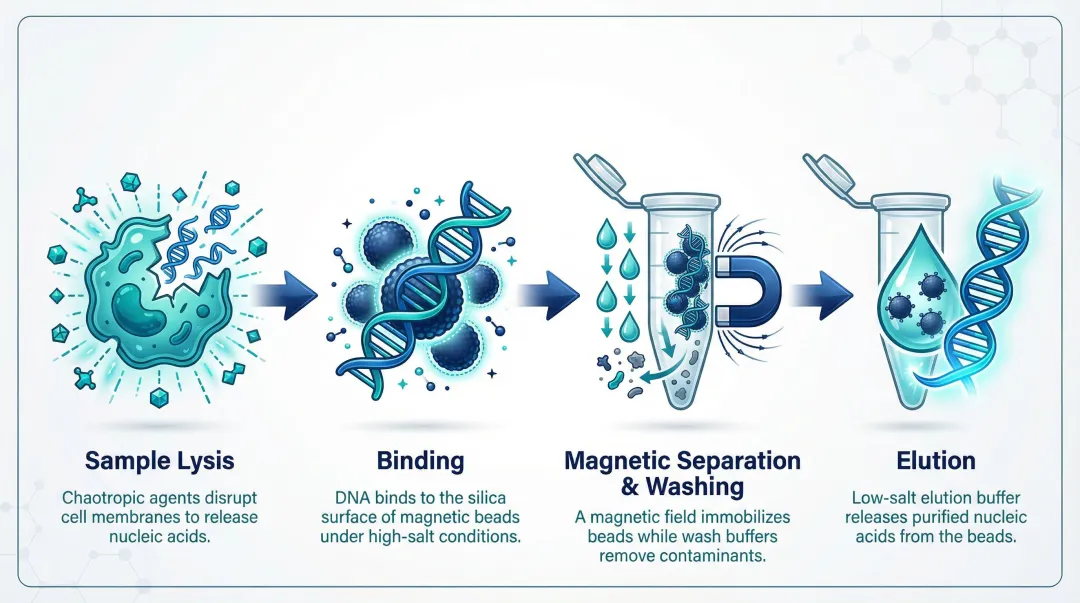
The process moves samples through four controlled phases: lysis, binding, washing, and elution. The magnetic field serves as the separation force—replacing centrifugation or gravity—so each phase can be precisely timed and automated.
Step 1: Sample Lysis
Chaotropic agents (such as guanidinium chloride or guanidinium isothiocyanate) perform lysis by simultaneously disrupting cell membranes, denaturing proteins, and inactivating nucleases. They also establish the high-salt, low-pH conditions required for nucleic acid binding to silica surfaces in the next step.
Lysis quality is the single most upstream variable determining extraction success. If cells are not fully disrupted, nucleic acids remain inaccessible to the beads regardless of binding chemistry efficiency.
Step 2: Binding
Silica-coated magnetic beads are added to the lysate and mixed. Under high-chaotropic-salt conditions, negatively charged nucleic acid phosphate backbones form hydrogen bonds and electrostatic interactions with the silica bead surface, selectively adsorbing DNA or RNA while proteins, lipids, and polysaccharides remain in solution.
Two bead properties directly shape extraction performance:
- Surface area — determines total binding capacity per reaction
- Coating chemistry — controls nucleic acid size preference and selectivity
Step 3: Magnetic Separation and Washing
An external magnet immobilizes bead-nucleic acid complexes against the tube or vessel wall within seconds, allowing the protein- and inhibitor-rich supernatant to be aspirated or poured off. Sequential wash buffers are then applied to remove residual chaotropic salts, ethanol, and co-precipitated contaminants without disrupting the bead-nucleic acid bond. The number and composition of wash steps directly impact downstream purity.
Step 4: Elution
Removing the magnetic field releases beads back into suspension. A low-salt aqueous buffer (or nuclease-free water) at neutral to slightly alkaline pH disrupts the surface interactions, releasing purified nucleic acid into solution. Elution volume, temperature, and incubation time can be adjusted to optimize concentration for specific downstream applications—including direct elution into barcoded tubes for clinical workflows.
Key Factors That Affect Extraction Quality and Yield
Bead Surface Chemistry and Particle Size
Silica-coated beads are the standard for total nucleic acid extraction. Specialized chemistries enable targeted applications:
- Carboxyl-modified beads allow size-selective enrichment—SPRI technology uses PEG and salt concentrations to bind DNA fragments selectively, with higher PEG ratios forcing smaller fragments to bind
- Streptavidin-coated beads target biotinylated nucleic acids
- CamSelect Long™ bead technology (Cambrian Bioworks' patented innovation) enriches high molecular weight DNA for long-read sequencing, increasing usable gigabases per flowcell by up to 3x
Critical Variables
Each of these variables requires deliberate optimization — a misstep at any stage compounds into downstream failures:
- Sample matrix complexity: High-lipid or high-protein samples (FFPE tissue, whole blood) need adjusted lysis volumes and duration. Hemolyzed samples introduce heme-based inhibitors requiring extra wash steps; cfDNA from plasma demands gentle lysis to avoid genomic DNA contamination.
- Lysis conditions: Chaotropic salt concentration, incubation time, and temperature must match the sample type. Under-lysis leaves nucleic acid trapped; excessive heat fragments it — RNA is especially vulnerable to thermal degradation.
- Bead-to-input ratio: Bead volume relative to nucleic acid load must stay within the validated range for each sample type. Too little leads to binding saturation and yield loss; too much promotes co-precipitation of non-target molecules.
- Wash stringency and step count: Insufficient washing carries inhibitors into the eluate, suppressing downstream PCR. Residual ethanol from over-washing is equally damaging. Two-phase wash protocols significantly improve qPCR reproducibility.
- Elution volume and temperature: Smaller elution volumes concentrate nucleic acid but can reduce total recovery. Warm elution buffers (50–70°C) improve recovery of high-molecular-weight DNA, while low-input samples like cfDNA benefit from reduced volumes to maintain detectable concentrations.
Common Misconceptions and Limitations
More Beads Does Not Equal Better Yield
Adding more beads doesn't improve yield—it often does the opposite. Excess beads increase co-precipitation of inhibitors and reduce purity. Bead-to-nucleic acid ratios must be optimized as part of protocol design; volume alone is not a reliable lever for better performance.
Confusing Bead Performance with Lysis Performance
If extraction yields are low, the root cause is often incomplete cell lysis rather than bead failure. The bead can only capture what has been released into solution—teams should validate lysis conditions before troubleshooting bead parameters.
When Bead-Based Extraction May Not Be Optimal
- Ultra-low-input samples: Without specialized low-bind protocols, bead surface adsorption losses can reduce recovery
- Long-fragment integrity: Spin column methods often perform better unless purpose-built long-fragment beads are used
- Bead aggregation risk: Improper storage or buffer incompatibility reduces separation efficiency—thorough vortexing or sonication before use resolves most cases
Conclusion
Magnetic bead-based separation works by leveraging surface chemistry for selective nucleic acid binding and magnetic force for rapid, clean separation—making it a fast, scalable, and automation-ready extraction method for clinical and research applications.
Extraction quality depends on understanding and controlling the full process—lysis, binding chemistry, wash stringency, and elution conditions—not just on the beads themselves. Labs that treat it as a black box risk inconsistent yields and downstream assay failures.
Labs implementing or upgrading to automated extraction can consider Manta by Cambrian Bioworks—a compact, CE-IVD marked instrument that runs magnetic bead-based DNA and RNA extraction in approximately 30 minutes. Trusted by 67+ labs across clinical and research settings, it is built for reproducibility and minimal hands-on time.
Frequently Asked Questions
How do magnetic beads bind to DNA and RNA during extraction?
During lysis, high-chaotropic-salt and low-pH conditions drive nucleic acid phosphate backbones to bind silica-coated bead surfaces through electrostatic and hydrogen bond interactions. Proteins and lipids stay in solution, enabling selective capture of DNA and RNA.
What is the difference between magnetic bead-based extraction and spin column extraction?
Both methods use silica chemistry for nucleic acid binding. Spin columns rely on centrifugal force through a fixed membrane, which limits automation and scalability. Magnetic beads are separated using a magnet, enabling full automation, variable throughput, and direct compatibility with liquid handling systems.
Can magnetic bead-based extraction be used for cell-free DNA isolation from plasma?
Yes, specialized magnetic bead protocols optimised for cfDNA use gentle lysis and low elution volumes to maximise recovery of highly fragmented, low-abundance DNA from plasma while minimising genomic DNA contamination from residual cells.
How long does magnetic bead-based nucleic acid extraction typically take?
Most magnetic bead-based extraction workflows complete in approximately 30–60 minutes from sample input to elution, with automated systems reducing hands-on time to near zero compared to manual spin column protocols.
What sample types are compatible with magnetic bead-based nucleic acid extraction?
Compatible sample types include whole blood, plasma, serum, nasopharyngeal swabs, saliva, urine, FFPE tissue sections, and cell cultures. Each sample type requires its own optimised protocol.
Can magnetic bead-based nucleic acid extraction be fully automated in a clinical lab?
Yes. Bead-based extraction is one of the most automation-friendly nucleic acid extraction formats. Because separation is driven by a magnetic field rather than a centrifuge or column, it integrates directly with automated liquid handlers and dedicated extraction instruments used in clinical labs.