Introduction
DNA extraction quality directly determines whether downstream assays—PCR, NGS, or sequencing—succeed or fail. Labs still lose time and sample quality with outdated phenol-chloroform methods or inconsistent spin-column workflows. Studies show that up to 90% of failed molecular oncology NGS tests stem from pre-analytical causes, specifically insufficient tissue availability or insufficient DNA yield. Poor extraction quality doesn't just waste samples—it wastes expensive sequencing runs, delays diagnoses, and forces time-consuming troubleshooting.
Magnetic bead purification addresses all three problems: it preserves high-molecular-weight DNA integrity, scales from single tubes to 96-well plates, and eliminates centrifugation-induced shear stress.
This guide covers the science behind magnetic bead chemistry, a validated step-by-step protocol, practical troubleshooting strategies, and a decision framework for selecting the right kit or automated system for your lab's sample types and throughput needs.
TLDR:
- Magnetic beads use PEG-mediated depletion attraction and salt-neutralized binding to reversibly capture DNA
- Bead ratio controls fragment size: 1.8× captures fragments >100 bp; 0.6–0.8× selects for HMW DNA
- Over-drying bead pellets reduces elution efficiency for fragments >10 kb
- Residual ethanol depresses A260/230 ratios and inhibits downstream enzymes
- Magnetic beads outperform spin columns in automation compatibility, HMW DNA preservation, and throughput scalability
How Magnetic Bead DNA Purification Works
The Core Mechanism: Superparamagnetism and Reversible Binding
Magnetic bead DNA purification relies on superparamagnetic beads—typically 1–5 µm polymer spheres with iron oxide cores coated with functional groups like silica or carboxyl residues. These beads magnetise only in an applied magnetic field. Once the field is removed, they retain no permanent magnetism—preventing spontaneous clumping and ensuring uniform resuspension between steps.
DNA binding occurs through two complementary physicochemical mechanisms:
PEG-induced depletion attraction (PEG 8000 at 16–20% w/v) acts as a molecular crowding agent, stripping water from DNA and forcing it from an extended helical coil into a compact globular state. This depletion attraction causes the collapsed DNA to precipitate and adsorb onto the bead surface.
Salt-mediated charge shielding works in parallel: high salt concentrations (2M NaCl or MgCl₂) neutralize the negative phosphate backbone of DNA and the charged bead surface. With electrostatic repulsion eliminated, the collapsed DNA binds tightly to the beads.
Bead Surface Chemistry and Selectivity
The surface chemistry of the bead determines how binding occurs—and critically, what size range of DNA fragments you can recover. Two chemistries dominate commercial products:
- Silica-coated beads bind DNA non-specifically under high-salt, low-pH conditions via hydrogen bonding between silica hydroxyl groups and DNA phosphates. These beads typically offer higher binding capacity.
- Carboxylated beads (SPRI beads) rely on PEG-mediated collapse in NaCl-rich buffers. The carboxyl groups provide electrostatic binding sites once charge repulsion is neutralized. These beads enable tunable size selection by adjusting PEG and NaCl concentrations.
The Four-Phase Magnetic Bead Workflow
All magnetic bead protocols share a universal structure:
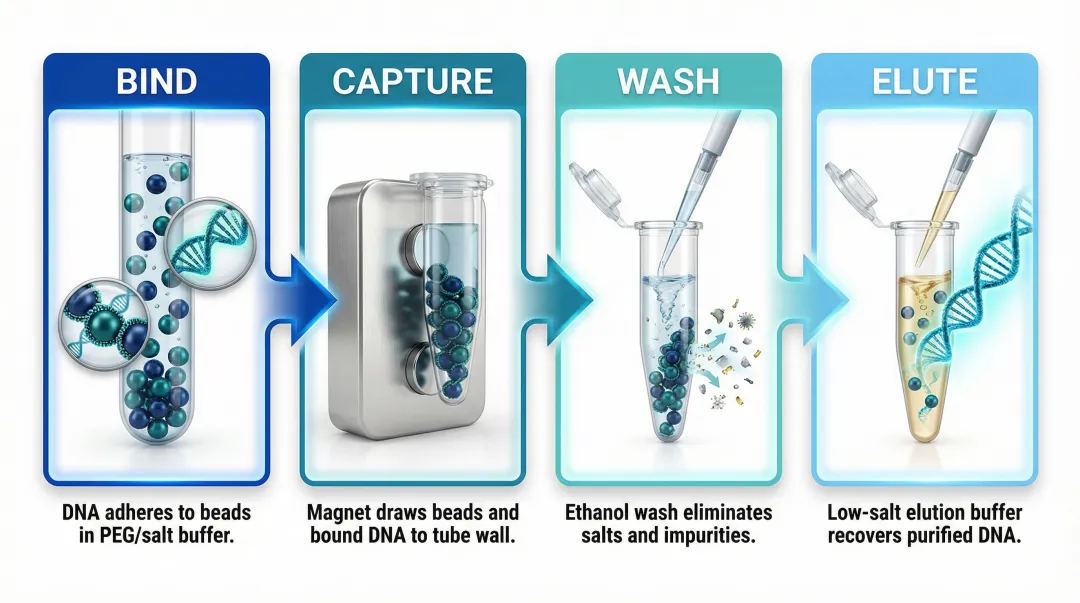
- Bind: DNA adsorbs to beads in binding buffer containing PEG and high-salt conditions. Mix thoroughly to maximise bead-DNA contact.
- Capture: Place the tube on a magnetic rack. Beads migrate to the wall within 1–2 minutes, leaving contaminants in the supernatant. Remove and discard the supernatant without disturbing the pellet.
- Wash: Add 70–80% ethanol wash buffer while the tube remains on the magnet. This removes residual salts, PEG, and enzyme inhibitors. Repeat 1–2 times depending on sample type.
- Elute: Remove the tube from the magnet and resuspend beads in low-salt elution buffer (10 mM Tris-HCl, pH 8.0) or nuclease-free water. A brief incubation at room temperature improves yield. Return to the magnet and transfer the purified DNA in the supernatant.