Introduction
Circulating cell-free DNA (cfDNA) forms the molecular foundation of liquid biopsy: a non-invasive approach for detecting cancer, fetal abnormalities, and transplant rejection. Unlike traditional tissue biopsies, cfDNA analysis relies on short DNA fragments (typically 130–170 base pairs) shed from apoptotic and necrotic cells into the bloodstream. These fragments circulate in plasma at extremely low concentrations, typically just a few nanograms per milliliter in healthy individuals.
While plasma is the preferred matrix for cfDNA isolation, extraction outcomes vary dramatically based on pre-analytical handling, extraction chemistry, and instrument choice. Research shows that 51% of published ctDNA studies fail to report the volume of blood from which plasma was extracted, and 42% omit plasma separation conditions — oversights that directly compromise assay reproducibility and clinical validity.
With cfDNA present at such scarce levels, even minor protocol deviations can push yields below the limit of detection for downstream assays.
This guide covers the complete extraction workflow, the critical variables that govern yield and purity, common pitfalls that sabotage results, and how to match your method to your specific liquid biopsy application. Whether your focus is circulating tumor DNA (ctDNA), non-invasive prenatal testing (NIPT), or transplant rejection monitoring, the principles here apply.
TL;DR
- Plasma is the gold-standard matrix for liquid biopsy — it contains fewer contaminating cells than serum and yields cleaner short-fragment DNA
- Extraction success hinges on four variables: collection tube choice, double-centrifugation protocol, cfDNA-optimized chemistry, and elution volume
- cfDNA is highly fragile and scarce — delays in processing, freeze-thaw cycles, and incorrect centrifugation speeds cause low or degraded yields
- Choose magnetic bead-based automation for reproducibility and throughput, or spin-column methods for higher yields in low-volume manual workflows
- Use fluorometric assays (such as Qubit) over spectrophotometric methods — they measure cfDNA accurately at low concentrations where absorbance-based tools fall short
What You Need Before Extracting cfDNA from Plasma
Every variable upstream of the extraction step directly controls how much intact cfDNA reaches your elution tube. This section covers the three preparation requirements—tube selection, equipment, and operator readiness—that determine whether your downstream assay succeeds.
Sample Collection and Tube Requirements
EDTA or citrate tubes are required for cfDNA work. Heparin tubes are strictly contraindicated because heparin directly interferes with real-time PCR and sequencing library preparation, inhibiting downstream analysis even if cfDNA extraction succeeds.
Standard EDTA tubes must be processed within 2–6 hours of draw to prevent leukocyte lysis and genomic DNA (gDNA) contamination. When EDTA blood sits unprocessed beyond this window, white blood cells rupture and release high-molecular-weight DNA into plasma, masking rare ctDNA mutations critical for cancer detection.
Cell-Free DNA BCT tubes (such as Streck or PAXgene) extend the processing window safely to 3–14 days by stabilizing nucleated blood cells and preventing gDNA release. Streck tubes maintain cfDNA stability at 6–37°C for up to 14 days, making them essential for labs with off-site collection or delayed processing schedules.
Equipment and Kit Requirements
Minimum equipment includes:
- Calibrated centrifuge capable of 1,600–1,900×g (swing-out rotor) and 16,000×g (fixed-angle rotor)
- cfDNA extraction kit validated for short-fragment recovery (50–200 bp)—either magnetic bead or spin-column chemistry
- Fluorometric quantification instrument such as Qubit for accurate low-concentration measurement
- Manual pipetting setup or automated liquid handler depending on throughput needs
Clinical labs requiring CE-IVD compliance also need an IVD-certified automated extractor. Cambrian Bioworks' Manta, for instance, completes cfDNA extraction in approximately 55 minutes with minimal gDNA contamination—meeting both speed and regulatory requirements for diagnostic workflows.
Skill and Compliance Readiness
Before starting extraction, confirm the following:
- Protocol distinction: cfDNA kits recover short, dilute fragments (50–200 bp) and cannot be substituted with standard blood DNA kits optimized for high-molecular-weight DNA
- Assay compatibility: Verify that NGS panels, ddPCR, or real-time PCR workflows accept your expected cfDNA input quantities and fragment sizes
- Input minimums: Many clinical NGS panels require 5–30 ng of cfDNA input, which sets your minimum plasma volume and extraction efficiency targets
How to Extract cfDNA from Plasma: Step-by-Step
This workflow applies to standard EDTA-collected whole blood processed within the recommended time window. Deviations at any step will affect final yield and purity.
Step 1: Plasma Separation (Double Centrifugation)
Two sequential spins are required to remove leukocytes, cellular debris, and apoptotic bodies — each a source of contaminating genomic DNA.
| Spin | Speed | Duration | Temp | Rotor | Purpose |
|---|---|---|---|---|---|
| 1st | 1,600–1,900×g | 10 min | RT | Swing-out | Remove blood cells; collect plasma layer |
| 2nd | 16,000×g | 10 min | 4°C | Fixed-angle | Remove residual debris and apoptotic bodies |
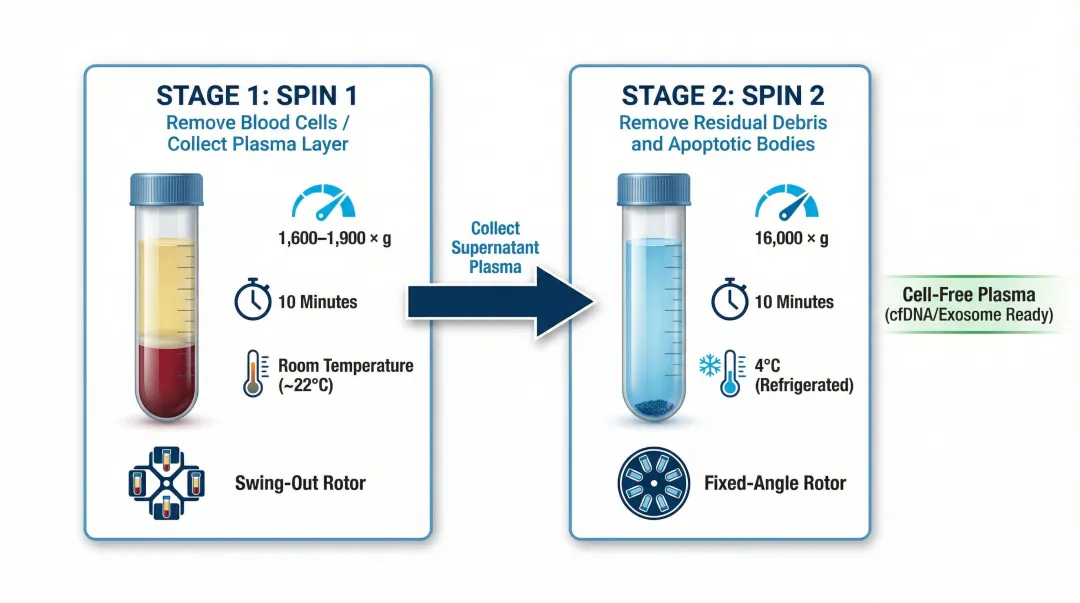
After the second spin, carefully transfer the clarified supernatant to a new tube, leaving the pellet undisturbed. Store at −80°C if extraction cannot proceed immediately.
Step 2: cfDNA Extraction (Binding Phase)
Use 1–4 mL of clarified plasma as input. Higher volumes directly improve total cfDNA yield and sensitivity for low-frequency variants — tests such as FoundationOne Liquid CDx and Guardant360 CDx use 4–8 mL specifically to detect variants with VAF below 0.5%.
Follow the manufacturer's lysis and binding protocol for your selected extraction chemistry:
- Spin-column methods bind DNA to silica membranes using centrifugal or vacuum force
- Magnetic bead methods capture DNA onto beads held in place by an external magnet
Ensure lysis buffer and proteinase K (where applicable) are equilibrated to the correct temperature before use. Incomplete lysis is a leading cause of low recovery.
Step 3: Washing
Perform all wash steps exactly as specified. Two failure modes are common here:
- Under-washing leaves PCR inhibitors that impair downstream assays
- Over-washing or excessive centrifugation force shears short cfDNA fragments off the membrane or beads
For magnetic bead protocols, confirm the magnet is fully engaged before aspirating wash buffer to avoid bead loss.
Step 4: Elution and Quantification
Elute in the smallest validated volume to maximise concentration (typically 20–50 µL for column methods; consult the kit manual for magnetic bead methods). Pre-warming the elution buffer to 55–70°C improves recovery of short cfDNA fragments that bind more tightly to silica membranes.
Quantify using a fluorometric method such as the Qubit dsDNA HS assay. Fluorometry is more accurate than spectrophotometric analysis (NanoDrop) at low cfDNA concentrations, because NanoDrop readings are skewed by co-eluted RNA, proteins, and other contaminants.
Verify fragment size distribution using a Bioanalyzer or TapeStation — particularly before NGS library preparation, where insert size directly affects library efficiency. Store eluted cfDNA in low-binding tubes at −20°C.
Key Parameters That Affect cfDNA Yield from Plasma
Even a validated protocol will produce inconsistent results if these variables are not actively controlled. Each parameter interacts with the others to determine final yield and quality.
The table below summarises the critical decision points across all four parameters:
| Parameter | Key Variable | Recommended Range | Primary Risk if Ignored |
|---|---|---|---|
| Centrifugation | Spin speed and number of spins | Double-spin; 2nd spin at 4,100–16,000×g | gDNA contamination from lysed cells |
| Input Volume | Plasma volume | 4–10 mL for ctDNA NGS panels | Falling below assay LOD |
| Extraction Method | Chemistry type | Match to downstream assay | Fragment size bias, yield inconsistency |
| Elution | Volume and buffer temperature | Pre-warm buffer; minimise volume for sensitivity | Loss of short fragments, over-dilution |
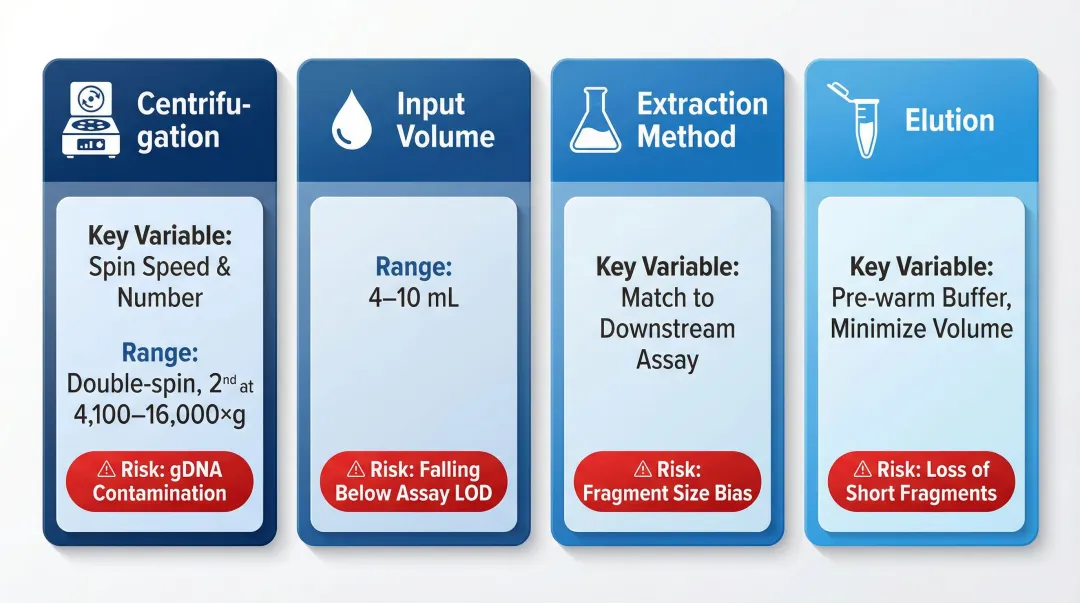
Centrifugation Speed and Protocol
Insufficient centrifugation leaves nucleated cells and platelets in the plasma. When these cells lyse during extraction or storage, they release high-molecular-weight genomic DNA that dilutes and interferes with cfDNA signal.
Studies comparing single versus double centrifugation demonstrate that double-spin protocols significantly reduce gDNA contamination, improving the sensitivity of ctDNA variant allele fraction detection in NGS and ddPCR assays. A second spin at 14,000–16,000×g removes remnant cellular debris that a first spin alone cannot eliminate.
Labs without high-speed centrifuges can use a validated alternative: 3,000–4,100×g for the second spin. Studies show no significant difference in ctDNA VAF when comparing 16,000×g to 4,100×g, which gives most clinical labs a workable option without specialised equipment.
Input Plasma Volume
On average, 1 mL of plasma contains approximately 2,000 genome equivalents of cfDNA. In healthy individuals, cfDNA concentrations typically range from a few nanograms per mL; disease states such as advanced cancer elevate levels, but baseline scarcity means total yield scales directly with input volume.
Larger input volumes increase total cfDNA mass but require kits with sufficient binding capacity. Using too little plasma risks falling below the limit of detection (LOD) of downstream assays. For ctDNA NGS panels with a median LOD of 1.25% VAF, recommended plasma volumes are 4–10 mL — a range that ensures adequate sensitivity for clinical decision-making. Getting centrifugation right in the previous step determines how much of that volume is clean, cell-free plasma rather than a contaminated fraction.
Extraction Method Chemistry (Spin Column vs. Magnetic Bead)
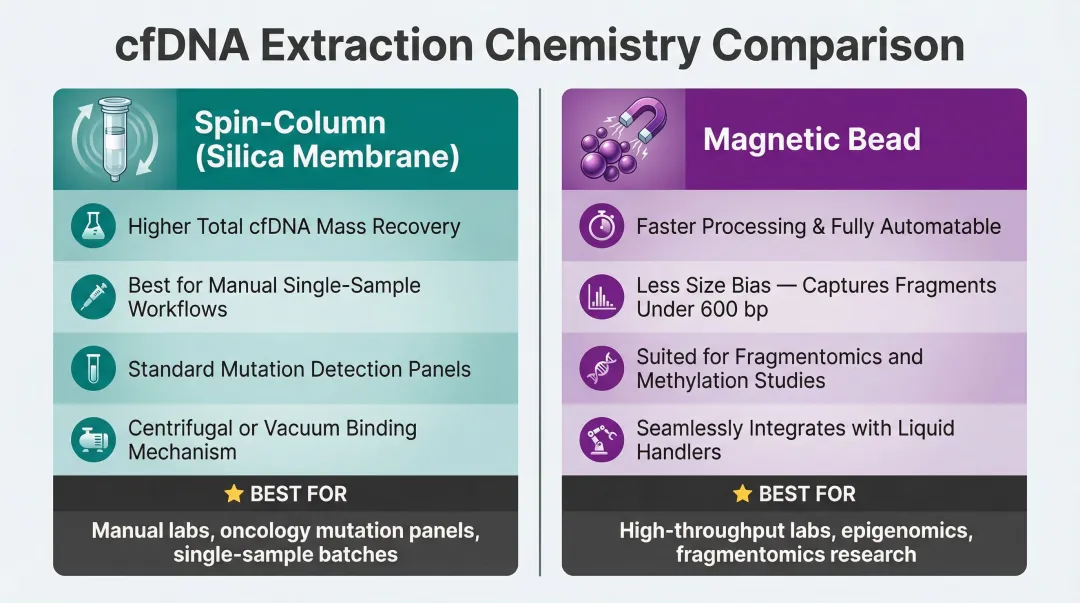
Silica-membrane spin-column methods (e.g., QIAamp CNA) typically recover higher total cfDNA mass for manual, single-sample workflows. Magnetic bead methods are faster, scalable, and more automatable — and they tend to be less biased against very short fragments (<600 bp), which matters when the assay targets highly fragmented ctDNA.
The right choice depends on what comes after extraction:
- Standard mutation detection panels: Spin-column methods offer reliable yield for low-throughput workflows
- Fragmentomics or methylation studies: Magnetic bead methods preserve short-fragment populations more consistently
- High-throughput or automated labs: Magnetic bead platforms integrate readily with liquid handlers, reducing hands-on time and operator variability

Multi-kit comparison studies confirm yield and reproducibility differences between method types, so validating kit performance against your specific assay before clinical deployment is not optional.
Elution Volume and Conditions
Eluting in a large volume dilutes cfDNA below detection thresholds; eluting in too small a volume may not fully recover DNA from the membrane or bead surface. The right balance depends on downstream assay sensitivity.
Concentration versus total recovery is a trade-off that must be optimised per application. Pre-warming elution buffer is important for very short cfDNA fragments (sub-100 bp) — these bind more tightly to silica membranes and require thermal assistance for efficient release. For bead-based systems, the elution buffer temperature and incubation time on-bead both influence recovery of this sub-100 bp fraction.
When Should You Choose Plasma for cfDNA Extraction?
Plasma is the preferred matrix for liquid biopsy work. Its low cellular content and compatibility with cfDNA-specific protocols make it the right starting point for most applications — but it's not the right choice for every scenario.
Primary use cases where plasma cfDNA extraction is the correct choice:
- Circulating tumor DNA (ctDNA) detection in oncology — tracking treatment response, minimal residual disease, and emerging resistance mutations
- Non-invasive prenatal testing (NIPT) for fetal aneuploidies, including trisomy 21, 18, and 13
- Donor-derived cfDNA monitoring in transplant medicine to assess graft rejection risk
- Infectious disease surveillance where viral or bacterial DNA circulates in plasma
- Pharmacogenetic profiling when whole blood is unavailable — genotyping concordance holds even at lower cfDNA yields
Not every workflow belongs on plasma, though. Here's when to consider an alternative matrix:
Scenarios where plasma is not the optimal choice:
- High-molecular-weight genomic DNA for germline variant calling or long-read sequencing — whole blood or buffy coat is the better source
- Limited sample volume where gDNA copy number matters more than cfDNA recovery
- Archived serum samples — serum carries higher background DNA from clotting-related cell lysis, making it a poor fit for ctDNA work, though retrospective studies have used it as a workaround
Common Mistakes and Troubleshooting When Extracting cfDNA from Plasma
cfDNA extraction failures are almost always traceable to specific, correctable errors in the pre-analytical or analytical phase.
Delayed or Improper Plasma Separation
Processing EDTA blood more than 2 hours after collection causes leukocyte lysis and massive gDNA release into plasma, masking rare ctDNA mutations and reducing assay sensitivity.
Fix: Use Streck BCT or process immediately. Always perform double centrifugation to remove cellular debris.
Using a Standard Genomic DNA Kit for cfDNA
Genomic DNA kits are optimised for large fragments (>10 kb) and will systematically lose cfDNA fragments in the 130–170 bp range during binding and washing steps.
Fix: Use kits validated specifically for short-fragment recovery. Published kit comparison data show yield differences of 5–20× in favour of cfDNA-optimised chemistries over standard genomic kits.
Skipping or Compressing Wash Steps
Residual ethanol or chaotropic salts from wash buffers inhibit PCR amplification and NGS ligation reactions, causing failed amplification despite adequate DNA input.
Solution: Complete all wash steps as specified. Include an optional dry spin before elution to remove residual ethanol.
Low Yield After Extraction
If cfDNA yield is consistently below expected range (healthy plasma: typically 5–20 ng/mL; cancer plasma: can reach 100–1,000 ng/mL), likely causes include:
- Input volume too low
- Incorrect kit for cfDNA (using a gDNA kit instead)
- Suboptimal elution temperature
- Excessive freeze-thaw cycles
What to do: Quantify with Qubit HS rather than NanoDrop, which overestimates cfDNA due to RNA co-elution.
Fragment Size Profile Appears Degraded
Yield problems address quantity — but quality matters just as much. If Bioanalyzer traces show smearing or loss of the characteristic ~167 bp mononucleosomal peak, the cause is likely DNase activity from delayed processing or thawing at an incorrect temperature.
Fix: Ensure plasma is aliquoted and frozen promptly. Thaw on ice before extraction to minimise degradation.
Frequently Asked Questions
Can you extract DNA from plasma?
Yes, plasma contains cell-free DNA (cfDNA) shed from apoptotic and necrotic cells, making it the preferred matrix for liquid biopsy applications. Plasma must be separated from blood cells using a validated double-centrifugation protocol and requires cfDNA-specific extraction kits to recover short, dilute fragments efficiently.
What is the best method for cfDNA extraction from plasma?
The best method depends on throughput and application. Spin-column silica-membrane kits tend to yield the highest total cfDNA mass for manual low-throughput workflows, while automated magnetic bead-based systems offer superior reproducibility and scalability for clinical labs processing multiple samples per run.
How do you minimise gDNA contamination during plasma cfDNA extraction?
Process blood promptly after draw using EDTA or BCT tubes, and perform a validated double-centrifugation protocol (1,600–1,900×g followed by 14,000–16,000×g) to deplete leukocytes and platelets before extraction. Skipping either centrifugation step is the most common cause of false signals in downstream ctDNA assays.
What is the cost of DNA extraction?
Costs vary widely by method. Manual cfDNA extraction kits typically range from a few hundred to several thousand rupees per sample depending on kit and input volume, while automated platforms carry higher upfront instrument costs but reduce per-sample cost and labour significantly at scale.
How much cfDNA is in 1 mL of plasma?
Whole blood contains far more DNA than plasma—genomic DNA from leukocytes is in the microgram range per mL. Plasma cfDNA is present at much lower concentrations, typically in the low ng/mL range in healthy individuals, though cancer patients can have significantly elevated levels.
Can you extract DNA from frozen plasma?
Yes, frozen plasma (stored at −80°C) is a valid source for cfDNA extraction and is widely used in retrospective studies. Multiple freeze-thaw cycles degrade cfDNA quality and should be minimised. Thawing on ice before extraction helps preserve fragment integrity.