Introduction
Magnetic bead-based nucleic acid extraction is a technique that uses superparamagnetic particles coated with nucleic acid-binding chemistry to selectively capture and purify DNA, RNA, or cfDNA from biological samples without centrifugation.
Lab managers and molecular scientists across clinical diagnostics, oncology, sequencing, and research labs run this method across hundreds of samples daily. Yet many struggle to explain why it works, what affects its performance, or when to use it over alternatives.
Automated magnetic bead systems process samples 74% faster than manual spin-column workflows. Even so, the underlying mechanism — how superparamagnetic iron oxide nanoparticles reversibly bind and release nucleic acids — remains poorly understood by many users.
This article breaks down the mechanism, the case for bead-based extraction over spin columns, how CamBeads™ technology implements it, and what drives extraction quality.
TL;DR:
- Superparamagnetic beads capture nucleic acids via surface chemistry (silica for DNA/RNA, carboxyl for cfDNA), then separate using an external magnet — no centrifugation needed
- Automated bead methods achieve 11.8% coefficient of variation vs. 16% for manual spin columns, delivering tighter reproducibility for clinical workflows
- Carboxyl-coated SPRI beads enable precise size selection for cfDNA workflows by altering PEG/salt ratios
- Bead size is a tradeoff: smaller beads bind more per mass, larger beads separate faster under magnetic force
- Bead-based extraction preserves fragile HMW DNA better than centrifugal shearing in spin columns
What Is Magnetic Bead-Based Nucleic Acid Extraction?
Magnetic bead-based extraction uses superparamagnetic iron oxide beads surface-coated with nucleic acid-binding ligands — silica for DNA/RNA, carboxyl groups for cfDNA. When an external magnetic field is applied, the beads and their bound nucleic acids migrate to the vessel wall, cleanly separating them from the rest of the sample matrix.
The result is a purified nucleic acid eluate, free from proteins, lipids, salts, and PCR inhibitors, ready for downstream applications like qPCR, ddPCR, or next-generation sequencing.
Unlike silica spin-column methods that rely on centrifugal force and vacuum filtration, magnetic bead extraction uses a magnetic field as the separation mechanism—removing the need for a centrifuge, making it inherently scalable and automation-compatible.
Key Advantages of Magnetic Separation:
- Eliminates centrifugation entirely, removing batch-processing bottlenecks
- Scales from 1 to 96 samples with no protocol changes
- Integrates directly with liquid handlers and robotic platforms
- Minimises mechanical shearing, protecting fragile DNA and RNA

Why Molecular Diagnostics Labs Rely on Magnetic Bead-Based Extraction
Throughput Without Batching
Magnetic bead extraction does not require batching in a centrifuge, so labs can process single samples or full plates on demand without waiting—critical for diagnostic labs with unpredictable inflow. Traditional spin-column workflows require accumulating samples to justify centrifuge runs, delaying time-sensitive results.
Clinical-Grade Reproducibility
Because all samples experience the same magnetic force, wash timing, and elution conditions (especially in automated formats), bead-based methods reduce operator-to-operator variability—a key requirement for clinical-grade reporting and IVD compliance. This consistency makes automated bead-based platforms well-suited for ISO 15189 and CAP-accredited labs where result reproducibility is non-negotiable.
Superior Recovery of Fragile Nucleic Acids
Low-input and fragile nucleic acids like cell-free DNA (cfDNA) from liquid biopsies are easily lost in centrifugation steps. Magnetic bead chemistry designed for cfDNA capture can recover these molecules with minimal carryover of high-molecular-weight genomic DNA. Magnetic bead methods preserve HMW DNA better than spin columns, which introduce mechanical shearing through centrifugal force.
Regulatory Alignment for Clinical Use
Clinical molecular labs are increasingly moving toward ISO 13485-compliant and CE-IVD workflows. Certified bead-based extraction platforms are now preferred over manual spin-column kits for regulated settings. Key regulatory requirements driving this shift include:
- CE-IVD marking under EU IVDR 2017/746 — mandatory for extraction kits used in clinical diagnostics
- Explicit differentiation from Research Use Only (RUO) products, which cannot be used for clinical reporting
- ISO 13485 quality management compliance for design and manufacturing traceability
Most spin-column kits are not certified for clinical IVD use, creating a compliance gap that bead-based platforms are specifically designed to close. CamBeads™ technology is built with exactly these regulatory requirements in mind.
How CamBeads™ Technology Works: Step-by-Step
Understanding how CamBeads™ work starts with the physics of their core material. Each bead contains a superparamagnetic iron oxide core — strongly attracted to a magnetic field when applied, but returning to a freely dispersed suspension the moment the field is removed. That reversibility is what enables automated workflows and prevents the irreversible clumping that limits other magnetic bead formats.
Cambrian Bioworks offers three surface chemistries, each designed for a specific extraction context:
- CamBeads-Si (silica-coated) — binds total DNA and RNA from high-input sample types like blood and tissue
- CamOxyl (carboxyl-coated) — optimized for cfDNA purification from low-input samples such as plasma
- CamSelect Long™ — a patented bead format engineered to enrich high-molecular-weight DNA for long-read sequencing workflows
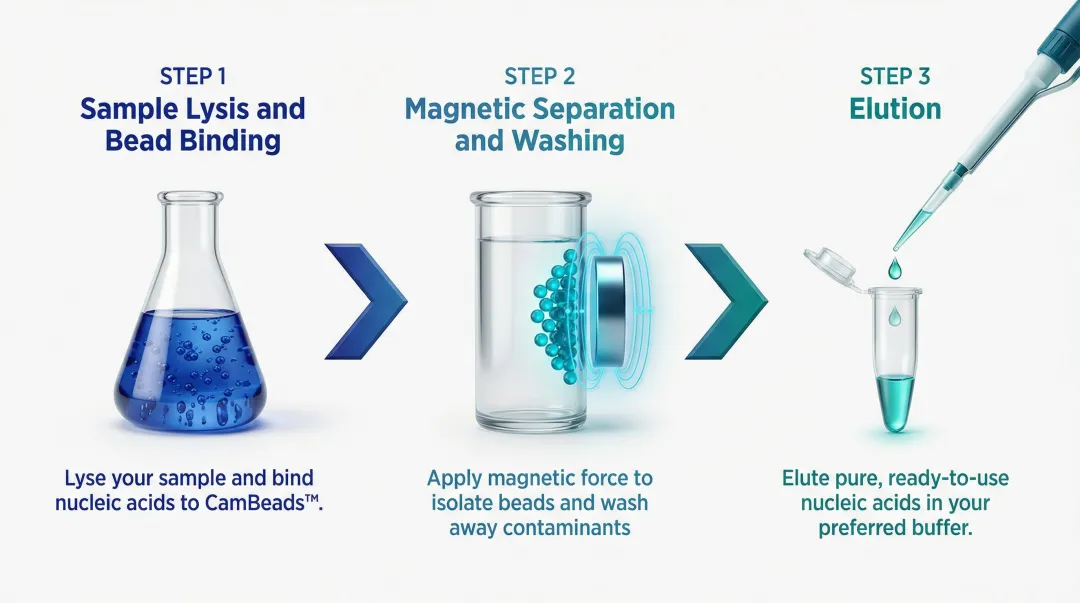
Step 1: Sample Lysis and Bead Binding
The biological sample (blood, plasma, tissue, swab, etc.) is lysed using a chaotropic salt buffer, which denatures proteins and releases nucleic acids. CamBeads™ are added to this lysate and nucleic acids adsorb onto the bead surface — driven by electrostatic interactions and hydrogen bonding between nucleic acid phosphate groups and the bead's silica or carboxyl surface chemistry.
Chaotropic salts like guanidinium thiocyanate disrupt hydrogen-bonding networks in water, causing nucleic acids to bind selectively to solid-phase surfaces rather than remain in solution — a principle well-documented in nucleic acid purification chemistry (Promega, DNA Purification Guide).
Step 2: Magnetic Separation and Washing
A magnetic field is applied (either via a magnetic rack or an automated instrument's integrated magnet), drawing the bead–nucleic acid complexes toward the vessel wall. The supernatant containing proteins, lipids, cell debris, and PCR inhibitors is removed. The beads are washed 2–3 times with ethanol-based wash buffers to remove residual contaminants while maintaining nucleic acid binding. The number and stringency of wash steps directly affect downstream purity ratios.
Step 3: Elution
The magnetic field is removed, allowing beads to resuspend. A low-salt elution buffer (or nuclease-free water) is added and mild heating disrupts the bead–nucleic acid interaction, releasing purified nucleic acids into solution. The eluate is then separated from the beads using the magnetic field one final time.
Elution volume is a key parameter: smaller volumes yield higher concentrations but may carry more residual inhibitors. As noted in the QIAamp DNA Blood Kit Handbook, eluting in volumes under 200 µL significantly increases final DNA concentration while slightly reducing overall yield — a trade-off labs should calibrate to their downstream assay requirements.
Key Factors That Affect Magnetic Bead Extraction Performance
Bead Chemistry and Size
- Silica-coated beads suit DNA/RNA from most sample types
- Carboxyl-coated beads are preferred for cfDNA due to their selectivity for short fragments
- Bead diameter affects surface area and capture kinetics — beads under 1 µm offer higher binding capacity per mass but slower magnetic separation; beads over 2 µm collect faster but with lower total capacity
Input Sample Quality and Matrix Composition
- FFPE-derived DNA is fragmented and cross-linked, requiring adjusted lysis conditions
- Plasma samples have high lipid content that can non-specifically bind beads
- Sample matrix composition directly affects binding efficiency: lipemic plasma causes volume displacement effects that alter the aqueous fraction available for bead binding
Automation Consistency
Manual magnetic racks expose beads to uneven magnetic force across the vessel, creating variable separation times and potential bead loss. Automated systems standardize magnet positioning, timing, and wash volumes, reducing the coefficient of variation across runs and delivering the reproducibility that downstream assays depend on.
Common Misconceptions About Magnetic Bead-Based Extraction
Myth: "Magnetic beads always co-purify genomic DNA when extracting cfDNA"
Bead chemistry (specifically carboxyl-coated beads at controlled pH and salt conditions) selectively binds short cfDNA fragments while leaving high-molecular-weight gDNA largely unbound. SPRI technology uses PEG and NaCl crowding agents to force nucleic acids out of solution. Increasing the ratio of SPRI reagent volume to sample volume increases binding efficiency for smaller fragments.
Properly designed workflows show minimal gDNA contamination. CamBeads™ cfDNA-specific bead formats leverage this selectivity for liquid biopsy applications.
Myth: "Spin columns give better purity"
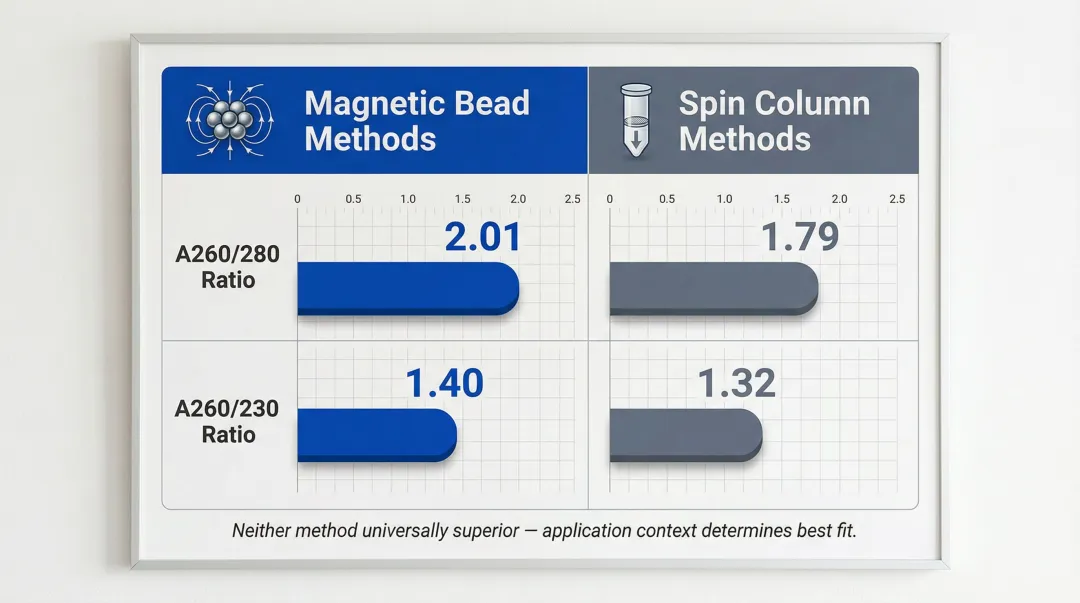
Purity (A260/280 and A260/230 ratios) depends more on wash buffer composition and technique than on the separation mechanism itself. In low-input and cfDNA applications, spin columns can introduce mechanical shearing and loss of fragile molecules that magnetic bead methods avoid. Magnetic bead methods show A260/280 ratios of 2.01 and A260/230 of 1.40, comparable to or better than spin-column results (1.79 and 1.32 respectively). Neither method is universally superior. Application context and bead chemistry choice determine what works best for your workflow.
When Magnetic Bead Extraction Is NOT the Best Choice
That said, magnetic beads aren't the right tool for every situation. For very high-input genomic DNA extractions where fragment length integrity is non-critical and throughput is low, phenol-chloroform or basic spin-column kits may be more cost-effective. Phenol-chloroform or silica spin-column protocols often yield the highest amount of DNA from challenging archival samples (such as museum specimens) compared to magnetic beads.
Magnetic bead methods add the most value when:
- Automation and hands-free processing are required
- Clinical compliance and audit trails are essential
- cfDNA recovery from low-input samples is the priority
- Consistent performance across diverse sample types is needed
Frequently Asked Questions
What are real-life applications of magnetic separation technology?
Industrial magnetic separation (mining, recycling) isolates ferromagnetic materials like iron ore from non-magnetic gangue. Biomedical applications use superparamagnetic beads coated with ligands or silica for nucleic acid extraction, immunomagnetic cell isolation, and protein purification—core techniques in molecular diagnostics and genomics.
What equipment is used in magnetic separation technology for nucleic acid extraction?
Key components include superparamagnetic beads (like CamBeads™), a magnetic rack or integrated magnet system, and sample tubes or well plates. Automated workflows add an instrument like Manta by Cambrian Bioworks, which handles magnet application, washing, and elution in a programmable sequence — CE-IVD certified for clinical DNA extraction.
What are examples of magnetic separation methods in life sciences?
Common methods include:
- Magnetic bead-based nucleic acid extraction (DNA, RNA, cfDNA)
- Immunomagnetic cell separation (such as circulating tumor cell capture)
- Magnetic bead-based protein purification
- SPRI (Solid Phase Reversible Immobilization) for NGS library clean-up
All rely on the same superparamagnetic principle but differ in bead surface chemistry.
What is the difference between magnetic bead extraction and spin-column extraction?
Spin columns use centrifugal force to push the sample through a silica membrane, while magnetic bead extraction uses an applied magnetic field to immobilize bead-bound nucleic acids. Bead-based methods are centrifuge-free, automation-friendly, and better suited for low-input and cfDNA samples where mechanical shearing must be minimized.
Can magnetic beads be used for RNA as well as DNA extraction?
Yes. Silica-coated magnetic beads bind both DNA and RNA under chaotropic conditions, making bead-based total RNA extraction standard for gene expression analysis and viral RNA detection. Some kits include DNase treatment steps or oligo-dT beads for selective mRNA capture.
What types of samples are compatible with magnetic bead-based nucleic acid extraction?
Compatible sample types include:
- Whole blood, plasma, serum
- Urine, saliva, swabs
- FFPE tissue, fresh/frozen tissue
- CSF (cerebrospinal fluid)
- Stool, cell cultures
Bead chemistry and lysis buffer require optimization per matrix; automated platforms manage multi-matrix workflows more consistently than manual methods.
Ready to optimize your nucleic acid extraction workflow? Cambrian Bioworks offers CamBeads™ magnetic bead technology alongside the Manta automated extraction platform—CE-IVD certified and trusted by 67+ labs globally. Learn more about CamBeads™ or explore how Manta's flexible 1-to-32 sample processing eliminates batching pressure in your lab.


