Introduction
Extracting both DNA and RNA from a single whole blood sample — without splitting it across two separate protocols — solves a core problem in molecular biology: sample scarcity. Simultaneous co-extraction isolates both nucleic acids in one workflow, cutting processing time in half and preserving limited material for downstream analysis.
This approach is particularly relevant when working with:
- Pediatric samples where draw volume is restricted
- Patients with hematological conditions requiring repeat testing
- Biobanked specimens with no possibility of recollection
- Studies requiring direct genotype-to-expression correlation from the same individual
This guide covers the methods available for co-extraction, how the protocol works, kit selection criteria, and when this approach is — and isn't — the right fit for your lab.
TL;DR
- Co-extraction isolates both DNA and RNA from a single whole blood sample, cutting required volume by ~50%
- TRIzol-based phase separation completes in ~2 hours and yields 3x more DNA than standalone methods
- NucleoSpin RNA + QIAamp DNA spin-column combinations yield high-purity results from frozen samples
- Thaw samples on aluminum blocks at room temperature — RNA integrity holds better than in 37°C water baths
- Quality benchmarks include A260/A280 ratios of 1.8–2.0 and RIN scores ≥7 for sensitive applications
Why Labs Choose Simultaneous DNA and RNA Extraction from Whole Blood
Resource Conservation and Sample Volume Constraints
The most immediate benefit of co-extraction is sample conservation. When blood volumes are limited—pediatric patients, longitudinal studies, or patients with hematological disorders—running separate extractions wastes precious material.
Co-extraction cuts the required volume roughly in half. Where traditional workflows might need 0.6 mL of whole blood (0.3 mL per extraction), co-extraction protocols recover both nucleic acid types from just 0.3 mL total, making every biospecimen go twice as far.
Scientific Value: Paired Genomic and Transcriptomic Analysis
Simultaneous access to DNA and RNA from the same individual enables direct correlation between genotype and transcriptional state—something sequential extractions from split samples cannot reliably deliver. Paired nucleic acid analysis has demonstrated clear diagnostic value across several clinical contexts:
- Splicing variants in cancer predisposition genes are detectable through paired germline DNA and RNA sequencing from whole blood—insights that DNA-only oncology panels routinely miss
- Integrated DNA/RNA profiling from EDTA whole blood characterizes mutations and expression patterns simultaneously, strengthening leukemia and lymphoma diagnostics
- Dual RNA isolation from sepsis samples simultaneously recovers host leukocyte RNA and bacterial RNA, opening direct host-pathogen interaction analysis from a single draw
Operational Efficiency and Cost Savings
Beyond sample conservation, co-extraction delivers measurable operational advantages. The TRIzol-based co-extraction protocol completes in approximately 2 hours, compared to 2 days required for traditional standalone phenol-chloroform DNA extraction, representing a 90% reduction in processing time. This time savings translates directly to reduced technician workload, lower reagent consumption per sample, and faster turnaround for urgent clinical cases.
Biobanking and Retrospective Research
That same efficiency advantage extends to archived collections. Biobanks and clinical research facilities need both DNA and RNA from stored samples—and re-collection is rarely an option. BBMRI-ERIC guidelines recommend EDTA tubes for whole blood intended for nucleic acid extraction, and co-extraction protocols validated for frozen EDTA blood allow retrospective studies on rare diseases and long-term outcomes without contacting patients again.
How Simultaneous Extraction Works: Methods, Protocol Steps, and Kits
Whole blood co-extraction functions by lysing blood cells to release nucleic acids, followed by separation and simultaneous or sequential recovery of DNA and RNA fractions. Each molecule requires different stabilization and precipitation chemistry, but optimized protocols process both concurrently to maximize efficiency.
Phase Separation (TRIzol-Based Co-Extraction)
The TRIzol method uses guanidinium thiocyanate and phenol to lyse cells and denature proteins. When chloroform is added and the sample is centrifuged, the homogenate separates into three distinct phases: a clear upper aqueous layer containing RNA, a white interphase containing DNA, and a red lower organic layer containing proteins. You then precipitate RNA and DNA concurrently from their respective fractions.
Modern optimized whole blood protocols complete in approximately 2 hours — dramatically faster than traditional sequential extraction.
Four Sequential Protocol Stages
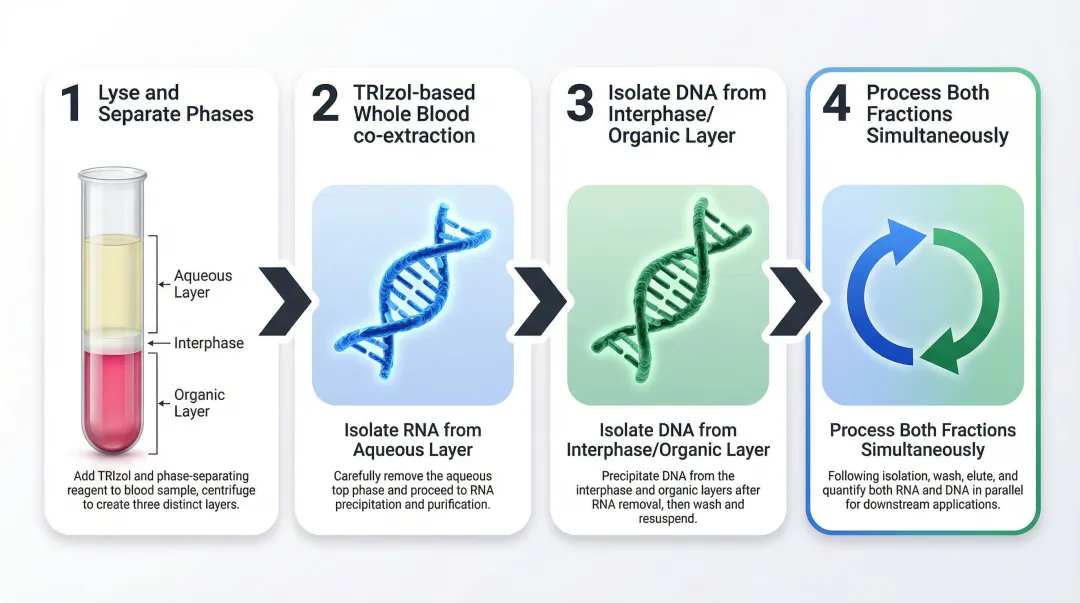
According to the TRIzol LS Reagent protocol, co-extraction follows four primary stages:
- Lyse and separate phases: Homogenize whole blood in TRIzol LS diluted 1:1 with RNase-free water, add chloroform, then centrifuge to create distinct aqueous, interphase, and organic layers
- Isolate RNA: Precipitate RNA from the aqueous phase using isopropanol, wash with ethanol, and resuspend in RNase-free water
- Isolate DNA: Precipitate DNA from the interphase and lower organic phase using absolute ethanol, wash, and resuspend in appropriate buffer
- Process both fractions simultaneously: The key to time savings is processing the RNA and DNA precipitation steps concurrently rather than sequentially
TRIzol co-extraction from coronary artery disease patient samples yielded DNA concentrations averaging 318 ng/µL — approximately 3-fold higher than standalone phenol-chloroform methods, demonstrating that co-extraction improves DNA recovery when protocols are optimized.
Spin Column Kit Combinations
NucleoSpin RNA Blood vs. PAXgene Performance
For frozen whole blood samples, dedicated spin-column kits offer an alternative to TRIzol-based methods. The Macherey-Nagel NucleoSpin RNA kit increased RNA yield by fivefold compared to the PAXgene Blood RNA Kit when extracting from frozen EDTA whole blood, making it a preferred choice for biobanked samples.
The complementary workflow typically involves:
- Extracting RNA first using NucleoSpin RNA Blood or Monarch Total RNA Miniprep kits
- Extracting DNA separately from the same thawed sample using QIAamp DNA Blood Mini Kit
- Both kits are validated for fresh or frozen whole blood stabilized with EDTA, citrate, or heparin
Critical Thawing Method Considerations
When working with frozen EDTA whole blood, thawing method significantly impacts yield and integrity. Thawing on an aluminum block at room temperature produced 20% higher DNA yield (25.19 µg/mL vs. 19.40 µg/mL) and better RIN scores (5.8 vs. 5.4) compared to rapid thawing in a 37°C water bath.
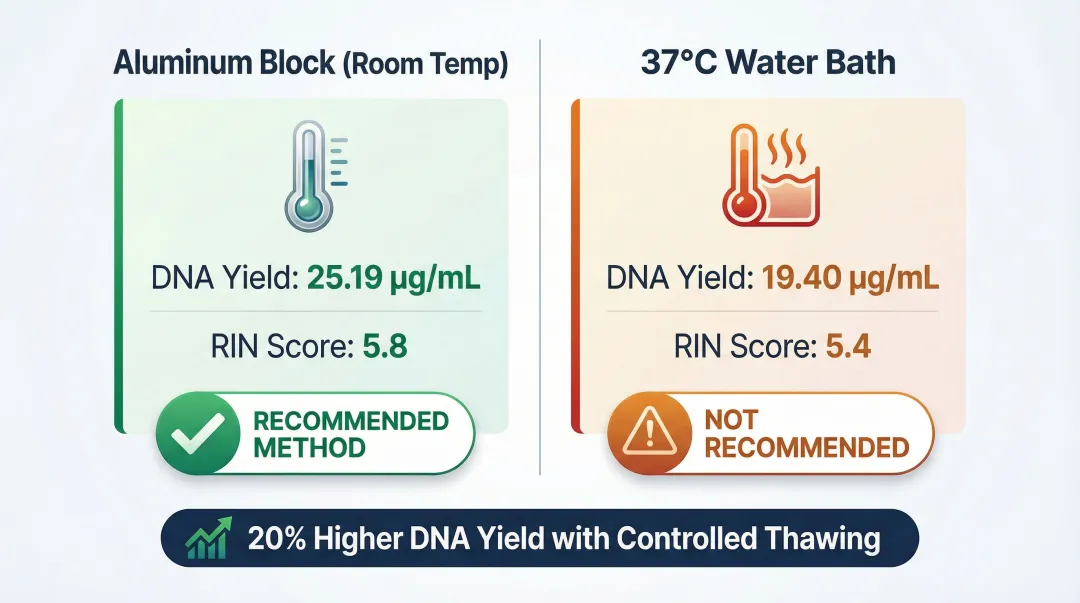

Slower, controlled temperature changes reduce RNase activation and minimize cell rupture. Rapid heating at 37°C activates sequestered RNases more aggressively, degrading RNA before stabilization reagents can act.
Automated Magnetic Bead-Based Extraction
Magnetic bead-based extraction offers a scalable alternative to manual methods for higher-throughput laboratories. Magnetic particles with selective surface chemistry bind nucleic acids, automated wash steps remove contaminants, and controlled elution recovers purified DNA or RNA — all without manual pipetting.
Platforms like Cambrian Bioworks' Manta bring this approach to clinical workflows. Key specifications relevant to whole blood processing include:
- Extraction time: ~30 minutes per whole blood run
- Throughput: No batching pressure — processes variable daily sample loads consistently
- Quality documentation: CE-IVD certified for clinical DNA extraction
- Reproducibility: Automated wash and elution steps eliminate operator-to-operator variability
Key Factors That Affect Co-Extraction Quality from Whole Blood
Blood Collection and Storage Conditions
EDTA tubes are the recommended standard for DNA/RNA co-extraction workflows. PAXgene tubes, while excellent for RNA stabilization, are explicitly not intended for the purification of genomic DNA from human whole blood according to manufacturer specifications.
For optimal results:
- Process fresh blood within 1 hour of collection when possible
- For long-term storage, freeze EDTA tubes at -80°C immediately after collection
- RNA and DNA of sufficient quality have been successfully extracted from frozen EDTA whole blood stored for up to 8.5 years when proper storage and thawing protocols are followed
RNase Control: The Critical Challenge
Whole blood leukocytes contain active RNases sequestered in cellular compartments. When freeze-thaw cycles disrupt these compartments, released RNases can almost completely degrade 28S and 18S rRNA, compromising RNA quality within minutes.
Essential RNase control best practices include:
- Use DEPC-treated or certified RNase-free water for all dilutions
- Use RNase-free plasticware and filter tips throughout
- Add RNA stabilization reagents immediately upon cell lysis
- Process samples on ice or at 4°C whenever possible
- Minimize freeze-thaw cycles—aliquot samples before freezing if multiple analyses are planned
Downstream Application Requirements
Quality thresholds vary by application:
| Application | Minimum A260/A280 | Minimum RIN | Notes |
|---|---|---|---|
| NGS library preparation | 1.8–2.0 | ≥7.0 | Illumina protocols recommend RIN ≥7 for RNA-seq |
| RT-qPCR | 1.8–2.0 | ≥5.0 | More tolerant of partial degradation |
| PCR genotyping | 1.7–2.0 | N/A | DNA only; moderate purity acceptable |
| Whole genome sequencing | 1.8–2.0 | N/A | DNA only; strict purity required |
| Microarray analysis | 1.8–2.0 | ≥7.0 | Similar to RNA-seq requirements |
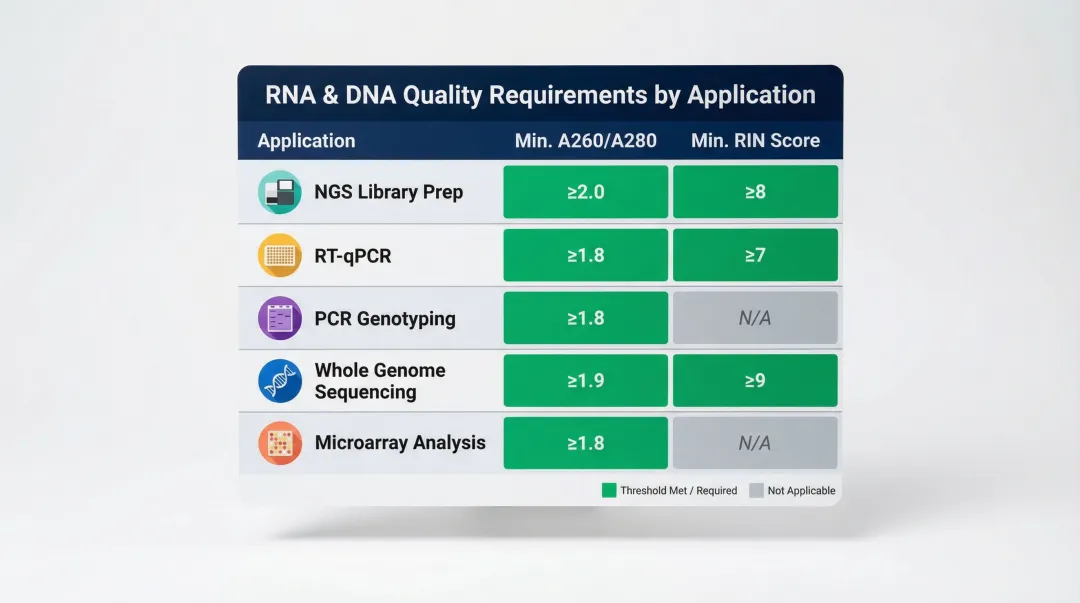

According to Thermo Fisher NanoDrop guidelines, A260/A280 ratios of ~1.8 indicate pure DNA, while ~2.0 indicates pure RNA. Ratios below 1.8 point to protein contamination; ratios above 2.0 in a DNA sample typically indicate residual RNA carryover.
Scale and Throughput Considerations
Maintaining those quality thresholds gets harder as sample numbers climb. Manual co-extraction handles 1–12 samples reliably, but at 50+ samples, concurrent RNA and DNA precipitation creates compounding pipetting errors and batch-to-batch variability that are difficult to control.
For routine high-throughput operations, automated platforms provide:
- Reproducible pipetting without operator fatigue
- Standardized protocols that reduce training requirements
- Sample tracking and audit trails for regulatory compliance
- Processing times of 30-60 minutes for 96-well plates
Common Mistakes and Misconceptions in Whole Blood Co-Extraction
Misconception: Co-Extraction Always Compromises Quality
Many labs default to separate extraction workflows, assuming co-extraction reduces purity or yield for one molecule. Many labs default to separate extraction workflows, assuming co-extraction reduces purity or yield for one molecule. In fact, optimized TRIzol co-extraction produced DNA yields 3-fold higher than standalone phenol-chloroform methods in published comparisons.
While DNA purity (A260/A280) may be slightly lower in co-extraction workflows, the difference is negligible for most downstream applications — PCR, qPCR, and RNA-seq all perform reliably with co-extracted material. Match the extraction method to your application, not to habit.
Incorrect Handling of Frozen Samples
Thawing at 37°C in a water bath is a common default in many labs, but this approach activates RNases more aggressively than controlled room-temperature thawing, leading to degraded RNA. Labs often blame the kit or protocol when the thawing step is the real culprit — a preventable error that wastes irreplaceable clinical samples.
Confusion Between RNA Stabilization and Extraction Compatibility
PAXgene tubes stabilize RNA in freshly collected blood but are not designed for high-yield genomic DNA extraction. Labs using PAXgene tubes expecting to also recover sufficient genomic DNA for whole genome sequencing will find yields insufficient. For workflows requiring both DNA and RNA, EDTA tubes paired with a validated co-extraction kit are the more reliable starting point.
Quick reference — common co-extraction errors:
- Thawing at 37°C: Accelerates RNase activity; thaw at room temperature instead
- Using PAXgene tubes for dual extraction: Optimized for RNA stabilization only; insufficient gDNA yield for WGS
- Assuming separate workflows are always superior: Co-extraction with optimized reagents can match or exceed standalone yields

When Simultaneous Co-Extraction May Not Be the Right Approach
Co-extraction isn't the right fit for every situation. Three specific scenarios call for a different approach.
When Maximum DNA Purity is Required
For applications demanding the highest DNA purity—such as whole genome sequencing with strict quality thresholds—a dedicated DNA extraction protocol will generally deliver better A260/A280 ratios. While TRIzol co-extraction yields high DNA quantities, standalone phenol-chloroform methods achieved significantly higher purity ratios (average 2.068 vs. 1.72). If only DNA is needed for your study, co-extraction introduces unnecessary complexity and reagent use.
When Sample Type is Incompatible
TRIzol-based co-extraction has been validated primarily for whole blood. Performance on clotted blood, plasma, serum, or other body fluids requires protocol modification — biological fluids with high contamination levels should be diluted 1:1 with RNase-free water prior to processing to prevent yield and purity losses.
Applying whole blood protocols directly to incompatible sample types is one of the most common sources of poor extraction results.
When Throughput is Too High for Manual Methods
Co-extraction protocols requiring concurrent manual precipitation of RNA and DNA from 50+ samples introduce compounding batch-level errors. At this scale, even experienced technicians face pipetting fatigue, timing drift, and sample tracking failures.
Automated extraction platforms provide the reproducibility and chain-of-custody traceability that manual workflows can't sustain at volume — a hard requirement for high-throughput clinical labs operating under accreditation standards.
Frequently Asked Questions
Can you extract both DNA and RNA from the same EDTA blood tube?
Yes. Both TRIzol-based phase separation and complementary spin-column kit combinations support co-extraction from EDTA tubes. EDTA is the preferred collection vessel for these workflows, preserving both nucleic acid types without the limitations of specialized stabilization tubes.
What is the best method for simultaneous DNA and RNA extraction from whole blood?
The best method depends on sample state and downstream use. TRIzol co-extraction is effective for fresh blood, offering time and cost advantages with completion in ~2 hours. For frozen samples, NucleoSpin RNA + QIAamp DNA kit combinations deliver better-validated performance with higher RNA yields and integrity.
How do you prevent RNA degradation during whole blood co-extraction?
Work on ice or at 4°C using RNase-free reagents and certified plasticware, and process samples immediately after collection when possible. For frozen samples, thaw on aluminum blocks at room temperature — not in a 37°C water bath — to limit RNase activation during lysis.
What A260/A280 ratio indicates good quality co-extracted nucleic acids?
An A260/A280 ratio of 1.8–2.0 is the accepted benchmark for DNA purity; RNA typically reads 2.0–2.2. Values below 1.8 suggest protein or phenol contamination, while values above 2.0 in DNA fractions may indicate RNA carryover or low-concentration samples affecting ratio accuracy.
Does freezing whole blood affect RNA quality during co-extraction?
Freeze-thaw cycles damage cell membranes and release RNases, accelerating RNA degradation. This can be mitigated by thawing on aluminum blocks at room temperature and using kits specifically validated for frozen blood rather than fresh-blood-only protocols.
Is simultaneous DNA and RNA co-extraction suitable for clinical diagnostic laboratories?
Yes, when validated kits and automated platforms are used. CE-IVD certified systems provide the traceability, reproducibility, and quality documentation required for diagnostic workflows. Manual methods typically need additional in-house validation before clinical deployment.