Introduction
Labs implementing liquid biopsy workflows face a recurring source of confusion: cfDNA and ctDNA appear across sequencing protocols, kit specifications, and clinical reports, yet are routinely treated as interchangeable. This overlap creates real operational problems — mismatched assay designs, compromised sample handling, and distorted result interpretation.
The distinction is not semantic. It directly affects laboratory performance metrics including sensitivity, specificity, and clinical utility.
Choosing the wrong analyte target or extraction approach compounds the problem downstream. Labs that conflate total cfDNA quantification with ctDNA profiling risk deploying incompatible workflows, consuming limited plasma samples, and generating unreliable data.
This guide breaks down the biological and operational differences between cfDNA and ctDNA — covering extraction requirements, workflow implications, and decision frameworks for lab professionals building or optimising liquid biopsy pipelines.
TL;DR
- cfDNA is all DNA circulating freely in plasma from any cell type — tumoral or non-tumoral
- ctDNA is the tumor-derived subset of cfDNA, carrying cancer-specific mutations and methylation signatures
- ctDNA is highly specific but represents less than 1% of total cfDNA in early-stage disease
- cfDNA quantification is cheaper and simpler; ctDNA detection requires sensitive assays like ddPCR or NGS
- Choose based on clinical goals — cfDNA for prognosis monitoring, ctDNA for mutation detection and companion diagnostics
What is cfDNA?
Cell-free DNA consists of double-stranded DNA fragments averaging 160–170 base pairs, released into blood plasma and other body fluids through normal cellular processes: apoptosis, necrosis, and active secretion. These fragments originate from multiple cell types including granulocytes, erythrocyte progenitors, lymphocytes, and hepatocytes. The modal fragment size of cfDNA distributes around 167 bp, corresponding to the nucleosomal unit wrapped around histone proteins.
In healthy individuals, plasma cfDNA concentrations typically range from 0–10 ng/mL (approximately 1,500 genome equivalents per mL). Cancer patients show substantially elevated levels (ranging from 0 to over 1,000 ng/mL) depending on tumor type and stage.
cfDNA elevation is not cancer-specific, though. Levels rise significantly in sepsis (averaging 41.2-fold higher than healthy controls), myocardial infarction, autoimmune disease, and even intense physical exercise. This non-specificity is the central laboratory challenge when using total cfDNA as a biomarker.
The half-life of cfDNA is short, ranging from 15 to 114 minutes in circulation. This rapid clearance enables real-time monitoring of disease states — but it also makes specimen integrity critical, since degradation begins the moment blood is drawn.
Plasma is the mandatory specimen type for cfDNA analysis. Serum contains 2- to 24-fold higher background genomic DNA than plasma, because leukocytes lyse during clotting and release genomic DNA. That contamination inflates cfDNA measurements and introduces noise that distorts downstream ctDNA detection assays.
Use Cases of cfDNA
Total cfDNA concentration — measured without mutation-specific analysis — provides clinical value in several applications:
- Non-invasive prenatal testing (NIPT) — detecting foetal chromosomal abnormalities
- Organ transplant rejection monitoring — measuring donor-derived cfDNA
- Infectious disease assessment — tracking pathogen-derived cell-free DNA
- Cancer prognosis — correlating elevated cfDNA with tumor burden
For oncology labs, total cfDNA concentration demonstrates prognostic value across multiple cancer types. In pancreatic cancer, pre-treatment cfDNA presence is associated with worse overall survival (HR = 2.42). In colorectal cancer, higher ALU-based cfDNA correlates with poorer outcomes (HR = 2.33). A cfDNA-based diagnostic model for pancreatic cancer achieved an AUC of 0.886 for distinguishing cancer from benign tumors, outperforming the CA19-9 protein marker (AUC 0.819).

The critical limitation: lack of standardisation. Collection tubes, extraction methods, and measurement protocols vary widely across studies, making cross-lab comparisons unreliable without protocol alignment. Extraction method variability alone creates up to 4.3-fold differences in DNA yield between commercial kits.
Labs investing in validated, automated extraction workflows reduce this variability. CE-IVD certified systems such as Cambrian Bioworks' Manta platform maintain consistency across runs by standardising the extraction process — which matters directly when total cfDNA concentration is driving clinical decisions.
What is ctDNA?
Circulating tumour DNA is the cancer-derived fraction of cfDNA, released specifically from tumour cells via apoptosis, necrosis, or active secretion. What distinguishes ctDNA from background cfDNA is both fragment size and molecular signature. ctDNA fragments are typically 20–40 bp shorter than nucleosomal cfDNA, with maximum enrichment between 90 and 150 bp. More importantly, ctDNA carries tumour-specific alterations: somatic mutations, copy number variations, structural rearrangements, and aberrant methylation patterns.
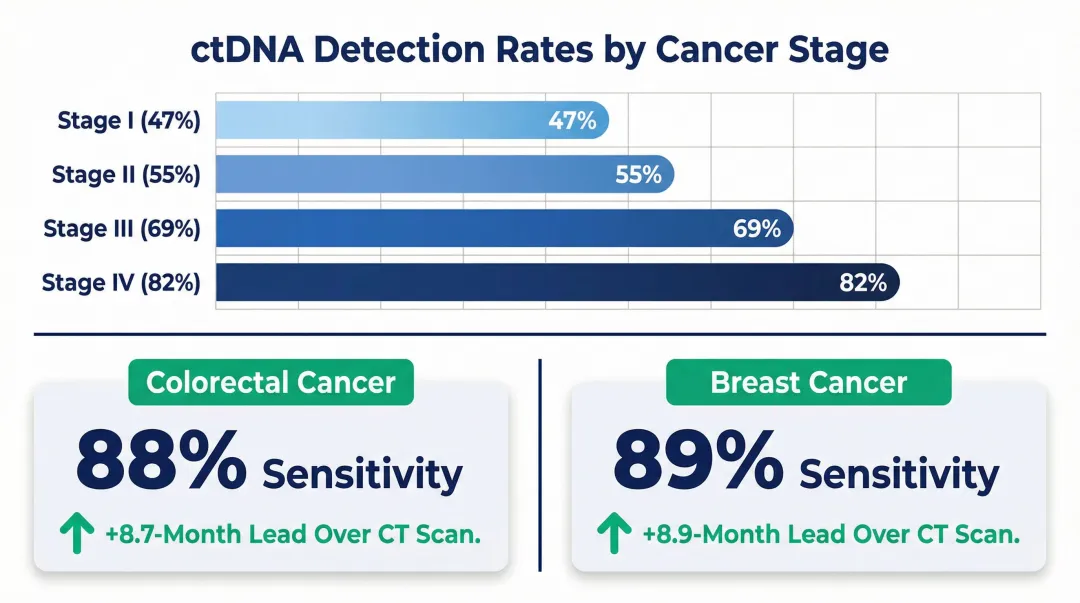
The ctDNA fraction problem is the defining challenge for liquid biopsy labs. ctDNA can represent anywhere from under 0.01% to over 90% of total cfDNA, depending on cancer type, stage, and tumour burden. In a landmark study, ctDNA was detected in 47% of stage I cancers, 55% of stage II, 69% of stage III, and 82% of stage IV cases. Early-stage, low-burden tumours shed very little ctDNA — which is why detection technology sensitivity is critical.
That low shedding directly shapes assay selection: labs must detect mutant allele frequencies as low as 0.001% to catch early-stage disease reliably. Digital PCR (ddPCR) achieves limits of detection of 0.001% VAF for BRAF V600E, while CAPP-Seq with integrated digital error suppression (iDES) reaches 0.0025% LOD.
Two main analytical strategies distinguish ctDNA from background cfDNA:
- Targeted mutation detection — using ddPCR, BEAMing, or targeted NGS (CAPP-Seq, TAm-Seq) when tumour mutations are known from tissue biopsy
- Methylation profiling — using bisulfite sequencing or enzymatic conversion methods to leverage tumour-specific epigenetic signatures without requiring prior mutation knowledge
Methylation approaches are most valuable for cancer-of-unknown primary cases and multi-cancer early detection screening, where mutation targets are not pre-specified.
Use Cases of ctDNA
ctDNA specificity is essential for applications requiring tumour-specific information:
Companion Diagnostics (FDA-Approved Tests):
- FoundationOne Liquid CDx (PMA P190032) — detects substitutions, indels in 311 genes, plus BRCA1/2 rearrangements and copy number losses
- Guardant360 CDx (PMA P200010) — identifies SNVs, indels in 55 genes, CNAs in 2 genes, and fusions in 4 genes
- cobas EGFR Mutation Test v2 (PMA P150044) — qualitative detection of defined EGFR mutations
Breakthrough Device Designations:
- Signatera (Natera) — personalised MRD assessment
- RaDaR (Inivata/NeoGenomics) — MRD detection in early-stage cancer
- Galleri (GRAIL) — multi-cancer early detection
Concordance data underscores ctDNA's clinical viability. Guardant360 CDx demonstrated 87.2% positive percent agreement and 99.9% negative percent agreement for panel-wide SNVs versus validated NGS tissue assays.
FoundationOne Liquid CDx showed 87.28% PPA and 99.83% NPA at the variant level for BRCA1/2 alterations — making ctDNA a viable non-invasive alternative when re-biopsy is not feasible.
For minimal residual disease monitoring, the lead-time advantage is well-documented. In colorectal cancer, serial ctDNA analysis identified relapse with 88% sensitivity and 98% specificity, with a median lead time of 8.7 months over CT imaging. In breast cancer, personalised ctDNA profiling detected metastatic relapse with 89% sensitivity and 100% specificity, with a median lead time of 8.9 months. These margins matter most when extraction quality is high enough to preserve the low-abundance fragments that carry the signal.
cfDNA vs ctDNA: Key Differences for Labs
| Feature | cfDNA | ctDNA |
|---|---|---|
| Origin | All cell types (tumoral + non-tumoral) | Tumour cells only |
| Fragment Size | ~160–170 bp (nucleosomal) | Enriched in 90–150 bp range |
| Concentration (Healthy) | 0–10 ng/mL | Not detectable |
| Concentration (Cancer) | 0–1,000+ ng/mL | 0.01%–90% of total cfDNA |
| Analytical Specificity | Low (non-specific to cancer) | High (tumour-specific) |
| Detection Method | Simple quantification (fluorometry, qPCR) | Sensitive mutation/methylation assays (ddPCR, NGS) |
| Cost Per Test | $15–50 | $200–1,500+ |
| Regulatory Maturity | Limited clinical use | FDA PMA approvals, Breakthrough Device designations |
Sensitivity and specificity tradeoffs define lab decision-making. Total cfDNA quantification is cheaper and simpler but lacks tumour specificity — elevated cfDNA can reflect inflammation, tissue damage, or exercise, not cancer. ctDNA is tumour-specific but requires low-error, high-sensitivity assays to detect.
Sensitivity variability by cancer stage is significant. In the GRAIL Galleri MCED validation study, sensitivity increased with stage: Stage I (16.8%), Stage II (40.4%), Stage III (77.0%), Stage IV (90.1%), with overall specificity of 99.5%. This stage-dependent performance reflects the ctDNA fraction problem: early-stage tumours shed minimal ctDNA.
That detection floor makes pre-analytical standardisation the operational bottleneck. When low-shedding samples are already at the limits of assay sensitivity, any upstream variability compounds the problem. The field still lacks consensus on:
- Collection tubes: EDTA vs. Streck Cell-Free DNA BCT vs. PAXgene ccfDNA
- Processing time: Acceptable window from collection to centrifugation
- Plasma separation: Protocol differences between labs
- Downstream handling: Freeze-thaw cycles and extraction kit selection
The tube choice alone has measurable consequences. Plasma DNA in EDTA tubes increases 10- to 20-fold after 4 days at room temperature due to leukocyte lysis, while Streck BCT and PAXgene ccfDNA tubes maintain stable concentrations over the same period. ESMO recommends cell preservation tubes to extend processing time from 4–6 hours (EDTA) to several days.
One common lab mistake is conflating cfDNA extraction kits with ctDNA assay performance. The extraction step determines how much total cfDNA enters the assay; the downstream sequencing or PCR step determines whether ctDNA within that cfDNA pool is detected. Both need independent validation.
Which Analyte Should Your Lab Prioritise?
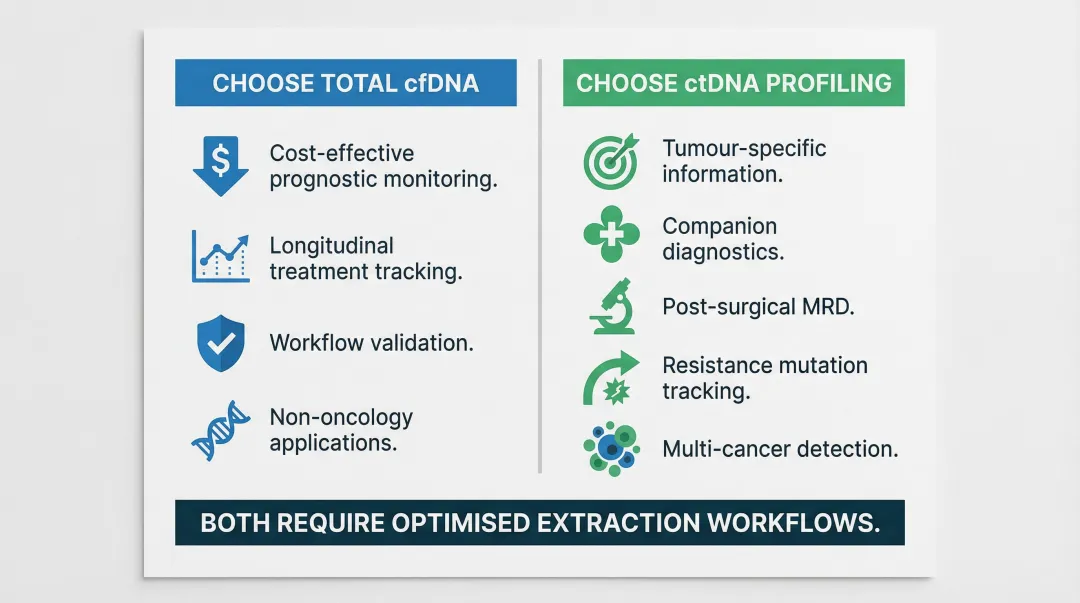
Choose total cfDNA concentration measurement when:
- The goal is cost-effective prognostic monitoring at scale
- Tracking treatment response longitudinally without known mutation targets
- Conducting early-stage workflow validation with limited resources
- Supporting non-oncology applications (NIPT, transplant rejection monitoring)
Choose ctDNA profiling when:
- The clinical goal requires tumour-specific information
- Running companion diagnostics for targeted therapy selection
- Performing MRD detection post-surgery
- Tracking therapy resistance mutations
- Conducting multi-cancer detection with tissue-of-origin determination
Whichever analyte you prioritise, the extraction workflow must be optimised for small fragments, low input volumes, and minimal genomic DNA contamination. Automated, magnetic bead-based systems reduce operator variability and improve inter-run reproducibility — critical when chasing ctDNA allele fractions below 1%. Cambrian Bioworks' Manta platform, which carries CE-IVD certification for clinical DNA extraction, is one example of this approach applied specifically to liquid biopsy workflows.

When neither analyte alone is sufficient, tiered approaches use both. A proposed two-step MCED screening framework uses low-cost protein/cfDNA tests for initial screening, followed by superior NGS-based tests for positive individuals. In a UK-based simulation of 5 million adults, this approach reduced false positives to 0.7% and lowered cost per cancer case detected from £172,828 (Galleri alone) to £33,534.
Conclusion
cfDNA and ctDNA are not competing methodologies. ctDNA is a biological subset of cfDNA. The right analyte choice depends entirely on the clinical question your lab needs to answer. Labs monitoring cancer prognosis at scale may find cfDNA quantification sufficient and cost-effective. Labs running companion diagnostics or MRD assays need the precision of ctDNA mutation profiling.
Getting the biology right starts before the sequencer. The most sensitive ctDNA assay will underperform if cfDNA extraction is inconsistent, pre-analytical conditions are unstandardised, or plasma inputs are contaminated with genomic DNA.
Labs that invest in validated, automated extraction workflows — and standardise collection protocols from tube type to processing time — position themselves to deliver reliable liquid biopsy results. Cambrian Bioworks builds purpose-designed automated extraction systems and cfDNA-specific reagent kits to help labs establish exactly this kind of consistent, scalable foundation for liquid biopsy.
Frequently Asked Questions
What is the difference between cell-free DNA (cfDNA) and circulating tumour DNA (ctDNA)?
cfDNA is all DNA circulating freely in plasma from any cell type — healthy or cancerous. ctDNA is specifically the tumour-derived fraction of cfDNA, carrying cancer-specific mutations or methylation changes. ctDNA fragments are typically shorter (90–150 bp) and present at very low concentrations in early-stage disease.
What is cell-free DNA (cfDNA)?
cfDNA consists of fragmented DNA (averaging ~160–170 bp) released into blood plasma through apoptosis and necrosis. It is present in all individuals — elevated in cancer but also in sepsis, myocardial infarction, and intense exercise, making it a non-specific marker requiring clinical context.
What is circulating tumour DNA (ctDNA)?
ctDNA is the cancer-specific subset of cfDNA shed by tumour cells, identifiable by somatic mutations, copy number variations, or tumour-specific methylation patterns. Its proportion within total cfDNA varies from less than 0.01% in early-stage cancer to over 50% in advanced disease.
How accurate is a circulating tumour DNA (ctDNA) test?
Accuracy depends on assay technology and cancer stage. Advanced platforms (ddPCR, CAPP-Seq) detect mutant allele frequencies as low as 0.001%, but clinical sensitivity for early-stage cancers remains limited (often under 50% for stage I). Specificity is typically very high (above 99% for most approved tests).
What do cell-free DNA (cfDNA) test results mean?
Elevated total cfDNA concentration indicates increased cell death associated with cancer progression, inflammation, or other pathology. Specific assays — such as NIPT or methylation-based tests like Epi proColon — interpret cfDNA characteristics in defined clinical contexts rather than relying on raw quantification alone.
What diseases are linked to circulating cell-free DNA (cfDNA)?
Beyond cancer diagnostics and prenatal testing, elevated cfDNA levels appear in sepsis, myocardial infarction, stroke, autoimmune diseases (lupus, rheumatoid arthritis), and organ transplant rejection.