Introduction
Picture this: An oncology lab receives a critical request to profile a patient's tumour for targeted therapy selection—but the only available sample is a biopsy block archived seven years ago. The DNA and RNA trapped inside this formalin-fixed, paraffin-embedded (FFPE) block hold the molecular answers the clinician needs, yet extracting usable nucleic acids from it remains unpredictable. Will the sample yield enough material? Will the data be reliable?
This uncertainty has a single root cause: formalin fixation preserves tissue architecture beautifully, but it chemically crosslinks and fragments DNA and RNA in ways that fresh-frozen samples never experience. FFPE samples are absolutely analysable—they are routinely used for FDA-approved companion diagnostics—provided your extraction protocol is built for degraded, crosslinked nucleic acids.
This guide walks you through what FFPE tissue is, why it remains the global standard despite its molecular trade-offs, and exactly how to extract DNA and RNA successfully from these samples using validated methods.
TLDR:
- FFPE = formalin-fixed, paraffin-embedded: tissue preserved in formaldehyde and encased in wax for long-term room-temperature storage
- FFPE blocks stored for 11–12 years yield DNA/RNA quality statistically identical to 1-year-old blocks
- Formalin crosslinks fragment nucleic acids (DNA: 100–400 bp; RNA: RIN 1–3), requiring FFPE-optimised extraction protocols
- Successful extraction demands deparaffinisation, heat-based crosslink reversal, and magnetic bead purification
- Standardising pre-analytical variables—fixation time and ischaemic time—is the single biggest lever for reproducible results
What Is FFPE Tissue and How Is It Prepared?
FFPE tissue is the result of a two-stage preservation process. Formalin-fixed means the tissue has been immersed in a 10% neutral-buffered formalin (NBF) solution—a formaldehyde-based fixative that halts cellular decay by crosslinking proteins, DNA, and RNA. This chemical reaction locks the tissue's molecular and structural features in place. Paraffin-embedded means the fixed tissue is then infiltrated with and encased in molten paraffin wax, which solidifies into a hard block that can be precisely sectioned into ultra-thin slices.
The resulting FFPE block is a solid wax cassette stored at room temperature in hospital pathology archives and biobanks. Because each block is linked to patient diagnosis, treatment history, and clinical outcomes, these collections form one of medicine's most valuable research resources—estimated at 400 million to over 1 billion samples worldwide.
Preparation: The Step-by-Step Process
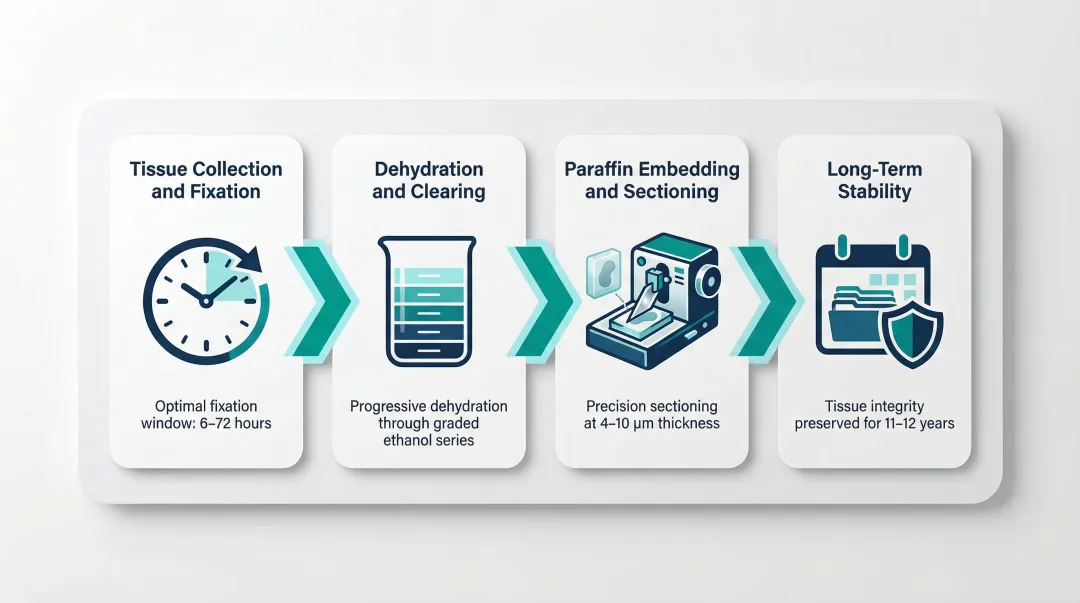
Tissue Collection and Fixation: Fresh tissue (biopsy or surgical excision) must be immersed in 10% NBF within 30–60 minutes of removal to minimise ischemic degradation. Fixation time should range from 6 to 72 hours depending on tissue thickness.
Under-fixation (<6 hours) compromises morphology and causes false-negative immunohistochemistry results. Over-fixation (>72 hours) introduces excessive crosslinking that impairs nucleic acid recovery and introduces sequencing artifacts.
Dehydration and Clearing: The fixed tissue is dehydrated through a graded ethanol series (70% → 100%) to remove water, since paraffin cannot penetrate aqueous tissue. A clearing agent—traditionally xylene, though safer alternatives like d-limonene exist—then displaces the ethanol, making the tissue ready for paraffin infiltration.
Paraffin Embedding and Sectioning: Cleared tissue is infiltrated with molten paraffin at approximately 60°C, then cooled into a solid block. A microtome cuts the block into sections: 4–5 µm for histology and immunohistochemistry, or 5–10 µm for nucleic acid extraction, where thicker sections maximise input material.
Long-Term Stability: Research shows no statistically significant difference in DNA, RNA, or protein quality between FFPE blocks stored for 11–12 years versus 1–2 years when processed consistently. This longevity is precisely what makes archived FFPE material usable for retrospective genomic studies—though extracting intact nucleic acids from it remains a distinct challenge.
Why FFPE Tissue Matters for Research and Diagnostics
Pathology departments worldwide have archived FFPE blocks for decades, spanning routine biopsies and surgical resections to samples collected during clinical trials. Each block carries documented patient history, making them irreplaceable for retrospective translational research that would be impossible to replicate prospectively.
Key Applications:
- Enables protein localisation and biomarker detection via immunohistochemistry (IHC) for disease diagnosis
- Supports somatic mutation detection, copy number variation (CNV) analysis, and gene fusion identification through next-generation sequencing (NGS)
- Identifies genetic variants that influence drug response in pharmacogenomics studies
- Tracks molecular changes across a patient's treatment journey using serially collected samples
These applications converge most visibly in precision oncology, where FFPE tissue powers FDA-approved companion diagnostics that directly guide therapy selection. FoundationOne CDx detects actionable alterations across 324 genes, microsatellite instability (MSI), and tumour mutational burden (TMB) using FFPE DNA. Oncomine Dx Target Test identifies mutations and fusions in 23 genes from FFPE DNA and RNA. These assays enable oncologists to match patients with targeted therapies, making FFPE the operational backbone of precision oncology.
FFPE Tissue vs. Frozen Tissue: Key Differences
Storage and Cost
Frozen tissue requires ultra-low temperature storage (−80°C or liquid nitrogen), demanding expensive freezers, continuous electricity, regular maintenance, and significant laboratory space. Large-scale frozen biobanking is logistically and financially prohibitive for most institutions. FFPE blocks, stored at room temperature indefinitely, cost a fraction to maintain—which is why clinical pathology defaults to FFPE.
Nucleic Acid Quality
Frozen tissue yields high-integrity nucleic acids: RNA Integrity Numbers (RIN) of 6–9 and high-molecular-weight DNA suitable for long-read sequencing. FFPE samples yield heavily fragmented material—RNA RIN values of 1–3 and DNA fragments typically 100–400 bp. However, FFPE-optimised NGS assays using short amplicons (<150 bp) achieve >95% coverage uniformity and >97% on-target reads, making most molecular applications feasible.
Morphology vs. Molecular Trade-Off
FFPE is unmatched for preserving tissue architecture and is the gold standard for histopathology and IHC. Frozen tissue is preferred when the highest-quality RNA or intact genomic DNA is essential. In clinical practice, FFPE is often the only available material—so extraction protocol optimisation is a practical necessity, not a workaround.
Here's how the two preservation methods compare at a glance:
| Parameter | FFPE Tissue | Frozen Tissue |
|---|---|---|
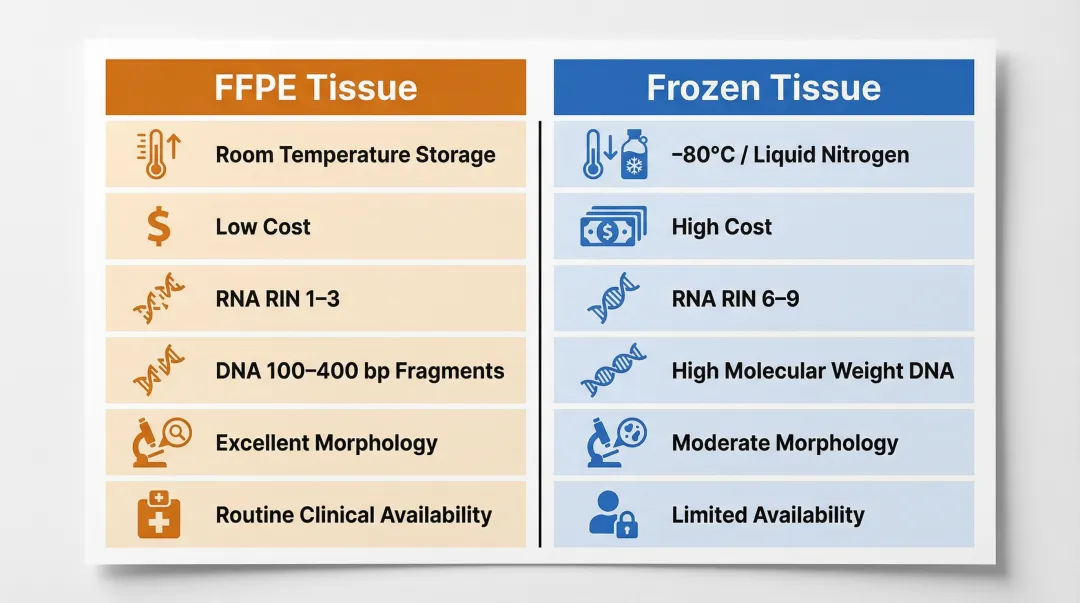
| Storage temperature | Room temperature | −80°C / liquid nitrogen |
| Storage cost | Low | High |
| RNA integrity (RIN) | 1–3 | 6–9 |
| DNA fragment size | 100–400 bp | High molecular weight |
| Tissue morphology | Excellent | Moderate |
| NGS compatibility | Yes (short-amplicon assays) | Yes (all assay types) |
| Clinical availability | Routine | Limited |
Storage and Cost
Frozen tissue requires ultra-low temperature storage (−80°C or liquid nitrogen), demanding expensive freezers, continuous electricity, regular maintenance, and significant laboratory space. Large-scale frozen biobanking is logistically and financially prohibitive for most institutions. FFPE blocks, stored at room temperature indefinitely, cost a fraction to maintain—which is why clinical pathology defaults to FFPE.
Nucleic Acid Quality
Frozen tissue yields high-integrity nucleic acids: RNA Integrity Numbers (RIN) of 6–9 and high-molecular-weight DNA suitable for long-read sequencing. FFPE samples yield heavily fragmented material—RNA RIN values of 1–3 and DNA fragments typically 100–400 bp. However, FFPE-optimised NGS assays using short amplicons (<150 bp) achieve >95% coverage uniformity and >97% on-target reads, making most molecular applications feasible.
Morphology vs. Molecular Trade-Off
FFPE is unmatched for preserving tissue architecture and is the gold standard for histopathology and IHC. Frozen tissue is preferred when the highest-quality RNA or intact genomic DNA is essential. In clinical practice, FFPE is often the only available material—so extraction protocol optimisation is a practical necessity, not a workaround.
Why Extracting DNA and RNA from FFPE Is Challenging
Formalin-Induced Crosslinking
Formaldehyde reacts with amino groups in nucleic acids and proteins, forming N-methylol adducts that condense into methylene bridges. This crosslinking traps DNA and RNA within a protein matrix, causing fragmentation during extraction. Formalin also inactivates DNA repair enzymes, letting cytosine deamination accumulate and creating artificial C>T or G>A transitions that masquerade as real mutations during sequencing.
RNA Is Disproportionately Affected
RNA's 2'-OH group makes it inherently vulnerable to hydrolysis. Combined with formalin modifications and the heat/chemical treatments used during tissue processing, RNA degrades significantly. Despite RIN values as low as 1.4, FFPE RNA remains usable for gene expression analysis when assays target amplicons ≤150 bp. The shift from RIN to DV200 (percentage of fragments >200 nucleotides) as a quality metric reflects this reality: DV200 >20% supports baseline NGS, while >50% enables optimal library yield.
Pre-Analytical Variables Amplify Challenges
Inconsistency in any of these parameters is the most common cause of extraction failure:
- Cold ischemia time: Delay between excision and fixation (target: <1 hour)
- Fixation duration: Under-fixation (<6 hours) or over-fixation (>72 hours) both compromise nucleic acid integrity
- Tissue thickness: Uneven formalin penetration in thick specimens
- Decalcification: Bone samples treated with strong acids suffer extreme degradation; EDTA is preferred
- Storage conditions: Temperature fluctuations over years
How to Extract DNA and RNA from FFPE Tissue: Step-by-Step
Deparaffinisation
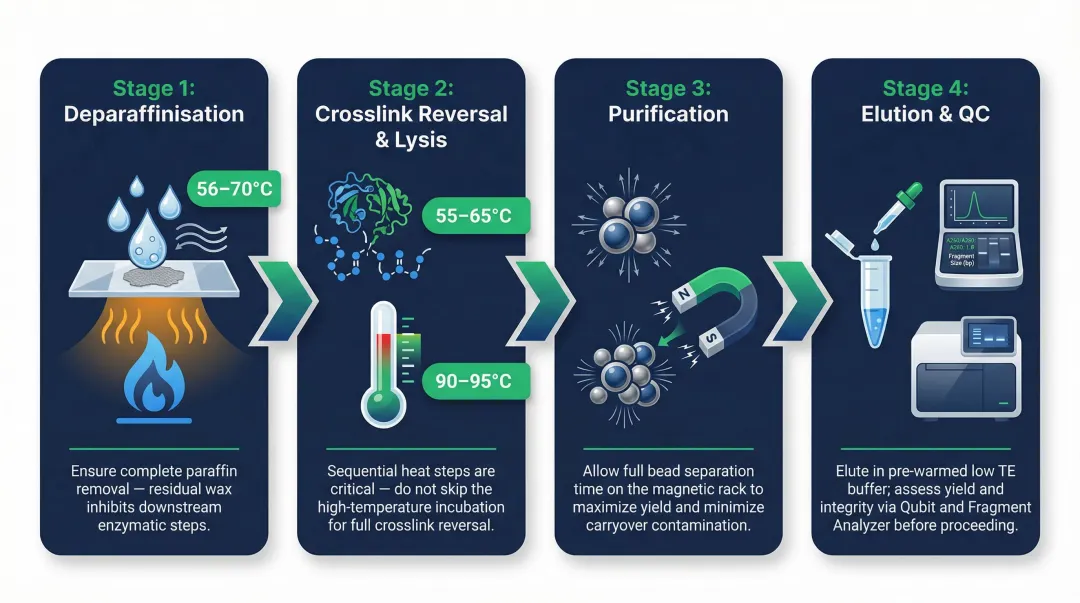
FFPE sections (typically 3–5 sections of 10 µm thickness) are heated at 56–70°C for 10–15 minutes to melt paraffin. The melted wax is dissolved using xylene (1–2 washes of 1 mL each), followed by ethanol washes (100%, 95%, 70%) to remove residual xylene. Xylene-free alternatives—mineral oil, d-limonene, or heat-based methods—yield equivalent DNA/RNA whilst eliminating toxic solvents and improving automation compatibility.
Crosslink Reversal and Tissue Lysis
After deparaffinisation, the tissue pellet is resuspended in lysis buffer and treated with Proteinase K at 55–65°C for 1–3 hours (or overnight). This enzymatic digestion breaks down the protein matrix and begins reversing formalin crosslinks. For RNA extraction, RNase-free reagents and lower incubation temperatures help preserve integrity.
A second, critical heating step completes crosslink reversal:
- Incubate at 90–95°C for 15–30 minutes to reverse residual formaldehyde-nucleic acid crosslinks — this improves RNA yield noticeably
- Under-incubation leaves crosslinks intact and reduces yield; over-incubation degrades already-fragmented nucleic acids — stay within the recommended window
Purification and Elution
Silica columns work adequately for DNA but can lose short fragments during washing. Magnetic bead-based purification recovers shorter DNA/RNA fragments more efficiently—critical for FFPE samples. Bead-based systems also scale better and integrate reliably with automated platforms.
For DNA, elute in low-salt buffer or water at 65–70°C to maximise recovery. For RNA, use RNase-free water and minimise room-temperature exposure post-elution. Dual-extraction kits enable co-extraction of DNA and RNA from the same FFPE section, conserving scarce biopsy material whilst enabling coherent multi-omics profiling.
Quality and Quantity Assessment
Once extraction is complete, QC assessment determines whether your nucleic acids are usable for downstream applications like NGS or PCR.
Quantification:Spectrophotometry (NanoDrop) overestimates FFPE nucleic acid concentrations by up to 30% because it cannot distinguish double-stranded DNA from single-stranded fragments and contaminants. Use fluorometric assays (Qubit dsDNA HS, RNA HS) for accurate low-concentration measurements before committing to NGS.
Integrity:Check 260/280 ratios (target ~1.8 for DNA, ~2.0 for RNA). Use Bioanalyzer or TapeStation to profile fragment size: FFPE DNA typically shows a smear of 100–500 bp; RNA RIN values of 1.4–3 are normal and workable. Run a test PCR amplifying a short amplicon (100–150 bp) as a functional quality check.
Best Practices to Maximise FFPE Extraction Yield and Quality
Standardise Pre-Analytical Variables
The single biggest driver of inconsistent results is variability in how tissue was originally processed. Where possible, use samples with documented fixation times (6–24 hours), known ischemic time (<1 hour), and consistent storage. For prospective collections, implement a standard operating procedure covering time-to-fixation, fixation duration, and storage temperature. Request fixation metadata when working with archival blocks.
Use FFPE-Optimised Kits and Assays
Generic nucleic acid extraction kits underperform with FFPE samples. Select reagents and assays matched to FFPE's specific challenges:
- Use kits validated for formalin-crosslinked, fragmented material
- For NGS, choose library prep kits designed for low-input, degraded DNA/RNA
- Design PCR primers to amplify amplicons <150 bp, staying within FFPE's typical fragment size range
Automate for Reproducibility and Throughput
Manual FFPE extraction introduces variability at every step (deparaffinisation, pipetting, incubation times, bead handling) that compounds across batches and between operators. Automated extraction platforms standardise each step, reduce hands-on time by up to 40%, and improve reproducibility across high sample volumes.
Cambrian Bioworks' Manta uses magnetic bead-based technology to deliver consistent FFPE DNA and RNA extraction with approximately 28-minute turnaround after lysis. It processes 1–32 samples per run, eliminating batching pressure in busy diagnostic labs. The system holds CE-IVD certification under EU IVDR and CDSCO IVD certification (MFG/IVD/2025/000069), making it suitable for regulated clinical settings across India and the EU.

Optimise Elution Volume and Storage
Post-extraction handling has a direct impact on downstream performance. Key recommendations:
- Elute in 30–50 µL to concentrate nucleic acids, but calibrate the volume to your specific downstream application to avoid material loss on membranes or beads
- Store FFPE DNA at -20°C and RNA at -80°C
- Aliquot samples immediately after extraction to avoid repeated freeze-thaw cycles
- Add RNase inhibitor to RNA elutions to preserve integrity during storage
Frequently Asked Questions
What does FFPE (formalin-fixed, paraffin-embedded) mean?
FFPE is a two-step tissue preservation method: tissue is first immersed in formalin, which crosslinks proteins and stabilises molecular structure, then embedded in paraffin wax to form a solid block. The resulting block is stored at room temperature and sectioned for downstream analysis.
What is an FFPE block?
An FFPE block is the solidified paraffin wax block containing embedded tissue. Pathology departments and biobanks store tissue in this format, cutting thin sections from it for microscopy, IHC, or nucleic acid extraction.
What is the purpose of paraffin embedding?
Paraffin embedding provides structural support to fixed tissue, enabling precise sectioning into thin slices (4–10 µm) using a microtome, whilst allowing room-temperature storage for decades without refrigeration.
What is deparaffinisation and what solvents are used?
Deparaffinisation removes paraffin wax from FFPE sections prior to extraction by heating to melt the wax, then treating with xylene and ethanol washes to clear residual solvent. Xylene-free reagents and commercial deparaffinisation kits are widely available as safer alternatives.
What thickness is recommended for FFPE sections?
For histology and IHC, sections are cut at 4–5 µm. For nucleic acid extraction, thicker sections of 5–10 µm are recommended, with multiple sections (3–5) pooled from a single block to maximise yield.
What is the difference between frozen tissue and FFPE?
Frozen tissue is snap-frozen and stored at -80°C or in liquid nitrogen, preserving higher-integrity nucleic acids at the cost of expensive cold-chain infrastructure. FFPE tissue trades nucleic acid quality for convenience — formalin crosslinking fragments DNA and RNA, but room-temperature storage makes it practical for long-term archiving.


