Introduction
FFPE tissue blocks are the backbone of cancer pathology, housing an estimated 400 million to over 1 billion archived tumor samples from patients worldwide. These specimens offer decades of clinical material — but extracting genomic data for NGS mutation analysis is far from straightforward. While FFPE remains the clinical standard for tissue preservation due to its ability to maintain cellular morphology, the formalin fixation process chemically damages DNA in ways that can silently compromise NGS results if not properly managed.
Formalin crosslinks DNA to proteins, fragments genomic material into short pieces, and introduces chemical modifications that generate false-positive variant calls. Each of these mechanisms compounds in the sequencing workflow.
When labs apply generic extraction protocols without accounting for these challenges, the consequences are compounding: failed library preparations, inflated artifact rates, unreliable biomarker calls, and ultimately, compromised treatment decisions for cancer patients.
Preventing those failures starts with understanding the biology — and enforcing the right quality gates before a single read is generated. This article covers the mechanisms behind FFPE DNA damage, the measurable quality standards labs must meet before sequencing, and a step-by-step workflow to consistently produce NGS-ready DNA from FFPE samples.
TLDR
- Formalin fixation fragments and chemically modifies DNA, making QC a prerequisite for reliable NGS from FFPE tissue
- Poor FFPE DNA quality inflates artifact rates by up to 20-fold, causing missed mutations and misclassified biomarkers like HRD and TMB
- Critical QC metrics — A260/280 (1.7–1.9), A260/230 (1.0–2.1), DIN score, fragment size, and functional yield — determine NGS readiness
- Structured extraction workflows with defined QC checkpoints cut typical FFPE failure rates by 25–40%
- CE-IVD certified platforms like Manta (Cambrian Bioworks) standardise FFPE extraction and reduce operator-dependent variability across runs
Why FFPE DNA Quality Is the Make-or-Break Factor in NGS Mutation Analysis
Formalin-fixed, paraffin-embedded (FFPE) tissue is the global standard for tumor biopsy preservation. Pathology archives worldwide hold an estimated 400 million to over 1 billion FFPE blocks, representing decades of clinical material. While FFPE preserves cellular morphology with high morphological fidelity for diagnostic pathology, the formalin fixation process creates chemical crosslinks between nucleotides and proteins, causing DNA fragmentation and chemical modification that complicates downstream genomic analysis.
The Molecular Consequences of Formalin Fixation
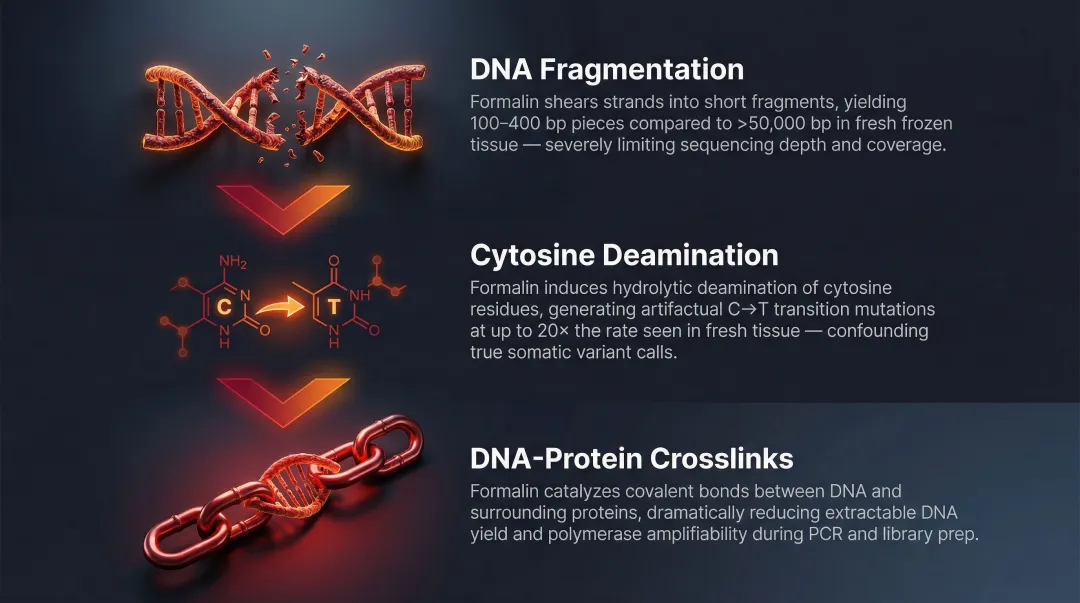
Formalin fixation dramatically alters DNA structure. DNA extracted from FFPE tissue is highly fragmented, showing high concentrations of small fragments approximately 100–400 bp in length — a stark contrast to fresh frozen tissue, where DNA fragments reach >50,000 bp. This fragmentation reduces library complexity, increases PCR duplicate reads, and lowers on-target coverage during sequencing.
Beyond fragmentation, formalin induces cytosine deamination, creating C→T transition artifacts. Whole genome sequencing of FFPE samples shows a median 20-fold enrichment in artifactual calls compared to matched fresh frozen samples, with a 7-fold specific increase in C>T/G>A transitions. These artifacts are not random — they systematically distort mutation profiles and confound variant calling algorithms.
Downstream NGS Impact
When FFPE DNA falls below quality thresholds, sequencing performance degrades measurably. Comparative WGS data illustrates the gap clearly:
| Metric | FFPE | Fresh Frozen |
|---|---|---|
| Mean sequencing depth | 77× | 93× |
| Proportion of regions at ≥70× depth | 0.351 | 0.782 |
Lower coverage reduces variant calling confidence and increases false negatives. Clinically relevant mutations can be missed entirely.
Clinical Stakes: Biomarker Misclassification
Poor FFPE DNA quality doesn't just reduce sequencing performance — it distorts clinically actionable biomarkers. FFPE-induced artifacts falsely elevate genome-wide Tumor Mutational Burden (TMB), showing a median TMB of 10.28 in FFPE versus 3.45 in matched fresh frozen tissue.
The consequences extend to Homologous Recombination Deficiency (HRD) detection. In matched cohorts, HRD scores fell below detection cutoffs for 7 out of 7 true HRD-positive cases using HRDetect, and 4 out of 7 cases using CHORD — resulting in incorrect classifications across the board.
When biomarkers like HRD, TMB, and microsatellite instability (MSI) are misclassified, patients may be excluded from targeted therapies or immunotherapies they would otherwise benefit from. This is why DNA extraction quality assessment must be treated as a clinical decision point, not a procedural checkbox — the integrity of every downstream result depends on it.
Key DNA Quality Metrics That Define NGS Readiness
A260/280 Ratio: Protein Contamination
This spectrophotometric measurement detects protein contamination. Pure DNA yields an A260/280 ratio of approximately 1.8 to 2.0, with acceptable NGS-grade DNA falling in the 1.7–2.1 range. FFPE-derived DNA typically achieves acceptable A260/280 values even when other quality dimensions fail — making this metric necessary but not sufficient for QC approval.
A260/230 Ratio: Organic and Salt Contamination
This ratio detects organic contaminants and chaotropic salts from extraction reagents. Pure DNA should yield A260/230 ratios of 2.0–2.2. FFPE-derived DNA commonly produces ratios of 1.0–1.3 due to residual contaminants from fixation and dewaxing steps. Severe organic contamination shows a spike at 230 nm and correlates directly with library preparation failure.
Low A260/230 values inhibit downstream enzymatic reactions — polymerases, ligases, and restriction enzymes all underperform, resulting in library prep failures and loss of limited sample material.
DNA Integrity Number (DIN) and Fragment Size Distribution
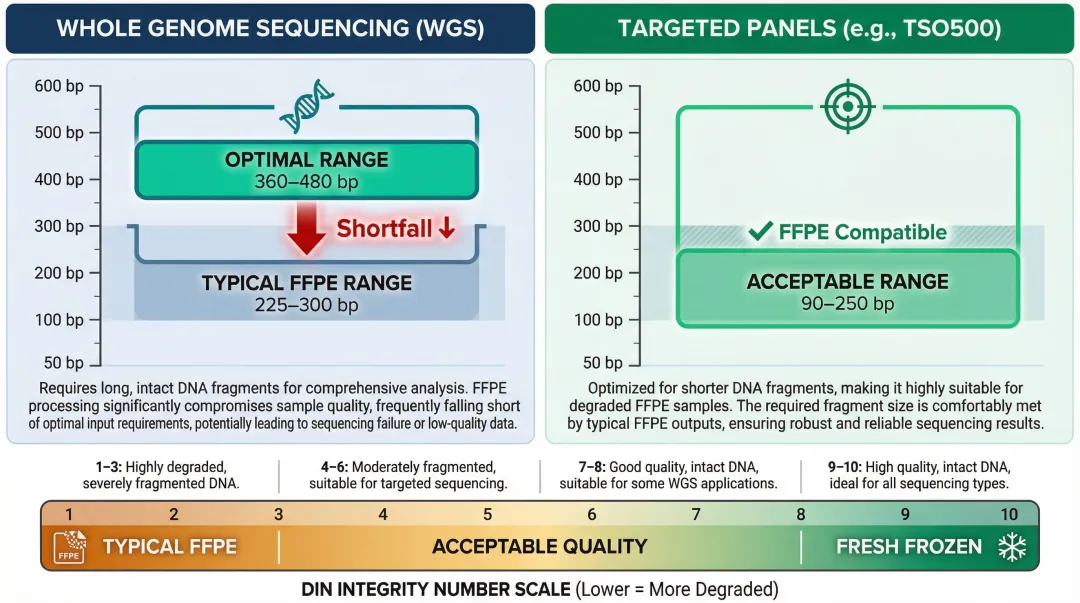
DIN, measured via platforms like Agilent TapeStation, provides a numeric score of overall DNA integrity on a 1–10 scale. FFPE-derived DNA routinely scores in the 1–3 range versus 9–10 for fresh frozen tissue. DIN score directly predicts fragment size distribution, and different NGS applications have different minimum fragment requirements:
- Whole genome sequencing (WGS): Optimal fragment size range is 360–480 bp; FFPE DNA often falls short at 225–300 bp
- Targeted panels: More tolerant of short fragments; Illumina TruSight Oncology 500 is optimized for 90–250 bp
Functional DNA Yield vs Total DNA Yield
Spectrophotometric quantification (NanoDrop) measures all nucleic acid, including degraded and non-amplifiable fragments. It significantly overestimates DNA concentration in FFPE samples — with a mean measured/expected ratio of 3.8 (SD = 6.4). Fluorometric quantification (Qubit) or qPCR-based functional DNA quantification is more predictive of NGS success.
Minimum input thresholds for common clinical NGS assays include:
- FoundationOne CDx: 55 ng
- Illumina TruSight Oncology 500: 40 ng
- Roche AVENIO Tumor Tissue Targeted Kit V2: ≥20 ng amplifiable DNA
Consistently meeting these thresholds starts with the extraction method. Manta by Cambrian Bioworks is CE-IVD certified for clinical DNA extraction and ISO 13485 certified, designed to deliver the purity and yield FFPE samples require — whether you're running a single sample or processing in volume.
The FFPE DNA Extraction Quality Workflow for NGS: Step by Step
The most common failures occur not from choosing the wrong kit, but from inconsistencies in pre-extraction tissue handling and insufficient mid-process QC. This workflow translates quality standards into practical checkpoints.
Step 1: Tissue Selection and Section Preparation
Select tumor-rich tissue sections from non-necrotic areas — necrotic tissue yields severely degraded DNA that will fail NGS QC regardless of extraction method. Standard sectioning practices include:
- Section thickness: 5–10 µm is preferable to single thicker sections (25–50 µm)
- Number of sections: FoundationOne CDx requires 10 unstained slides at 4–5 microns thick to achieve 1 mm³ tissue volume
- Macrodissection: Pathologist-guided dissection improves tumor content and downstream variant allele frequency accuracy
Step 2: Deparaffinization
Complete paraffin removal is essential — residual paraffin inhibits protease and polymerase activity in downstream steps. Options include:
- Xylene-based deparaffinization: Traditional method, effective but requires fume hood
- Heat-based deparaffinization: Preferred in clinical workflows for safety and consistency; alternatives include mineral oil, hexadecane, or tetradecane
Inadequate deparaffinization is a common hidden cause of low yield and impure extracts.
Step 3: Tissue Lysis and Formalin Crosslink Reversal
Proteinase K digestion at controlled temperature (typically 55–70°C) both lyses cells and reverses formalin-induced DNA-protein crosslinks. A validated protocol utilises 40 µl of proteinase K for 24 hours at 56°C, followed by 1 hour at 90°C for crosslink reversal.
Incubation time and temperature directly determine outcome:
- Under-digestion: Leaves DNA bound to protein, reducing yield
- Over-processing: Further fragments already damaged DNA
- Protocol deviations: Reducing proteinase K volume from 40 µl to 20 µl caused 12% of samples to fail, yielding no DNA at all
Standardised protocols consistently outperform improvised manual methods.
Step 4: DNA Isolation and Purification
Two dominant purification approaches:
- Silica membrane-based: Simple but can lose short DNA fragments and retain inhibitors; may underestimate residual DNA by up to tenfold
- Magnetic bead-based: Superior recovery of fragmented DNA and lower inhibitor carryover; automated magnetic bead extraction presents optimal outcomes and faster performance
For FFPE specifically, magnetic bead-based recovery matters more than it does with high-quality fresh frozen DNA. Short, damaged fragments are easily lost by silica membranes — and those fragments often carry the variant signal you're trying to detect.
Cambrian Bioworks' Manta uses magnetic bead-based extraction and completes FFPE DNA isolation in approximately 28 minutes after lysis. It processes 1 to 32 samples per run without batching pressure and delivers DNA integrity numbers (DIN ~5) suitable for cancer sequencing workflows.
Step 5: Quality Assessment (QC Gate)
This is the critical decision checkpoint. Measure:
- A260/280 (acceptable: 1.7–1.9)
- A260/230 (acceptable: 1.0–2.1 for FFPE)
- Fragment size via TapeStation or Bioanalyzer (DIN ≥2 preferred for most NGS)
- Functional yield via fluorometric or qPCR methods (minimum 20–55 ng depending on assay)
What to do when samples are borderline:
- Re-extract with adjusted protocol
- Use a lower-input library prep kit
- Flag sample for pathology review
- Never send sub-threshold DNA forward to sequencing
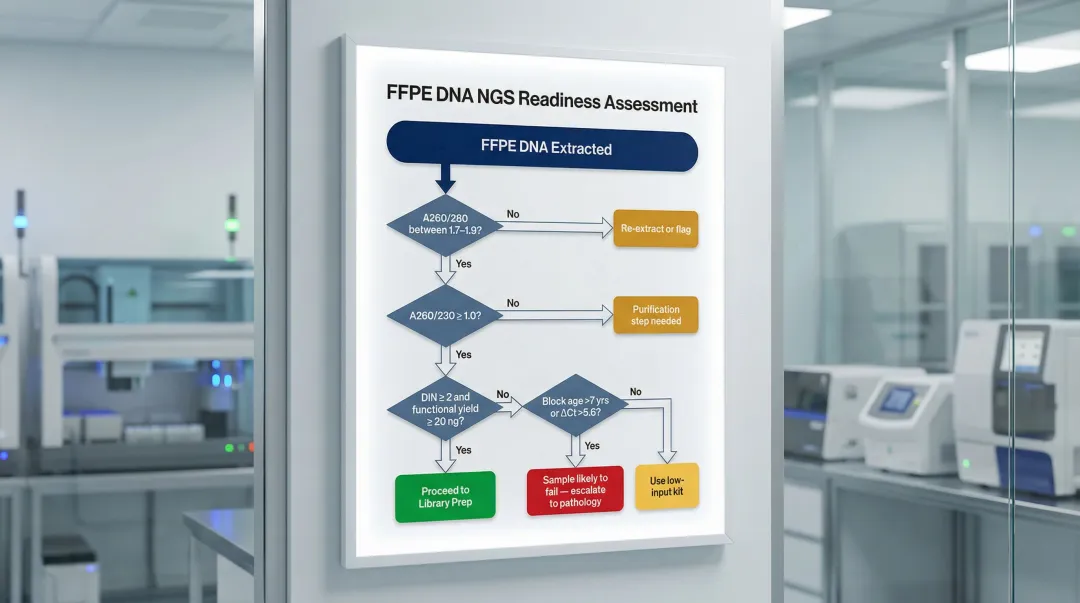
Decision trees for FFPE DNA amenability to NGS utilise combinatorial thresholds: >7 years of block storage, Illumina FFPE QC kit ΔCt > 5.6, and DIN < 2.0 indicate samples highly likely to fail.
Step 6: Library Preparation Decision and Handoff
QC-passed FFPE DNA proceeds to library preparation. FFPE-specific library kits differ from fresh frozen protocols in three important ways:
- Account for short insert sizes (100–600 bp)
- May require higher input amounts (40–55 ng vs 10 ng)
- Include steps to repair oxidative and deamination damage: uracil-DNA glycosylase (UDG) treatment dramatically reduces C:G > T:A artifacts
UDG pretreatment is particularly effective for severely fragmented FFPE DNA with qPCR ΔCt ≥1.55. Skipping FFPE-optimised library prep after DNA QC is the last place variant calling can be silently compromised — and one of the most avoidable failures in the workflow.
Common Mistakes That Cause FFPE DNA to Fail NGS Quality Gates
Three failure patterns account for the majority of FFPE samples that don't pass NGS quality gates — and each is preventable with the right protocols in place.
Relying Solely on Nanodrop for DNA Quantification
A260/280 and A260/230 readings from Nanodrop can appear acceptable even when DNA is severely fragmented or contaminated with PCR inhibitors. Labs that skip fragment size analysis and fluorometric quantification frequently send compromised DNA forward, resulting in library prep failure or inflated artifact rates in sequencing data.
Using Tissue Sections with Excessive Necrosis, Low Tumor Cellularity, or Poorly Fixed Areas
Pre-analytical variables are frequently underestimated:
- Fixation time: ASCO and CAP guidelines mandate 6–72 hours in 10% neutral buffered formalin. Too short = incomplete crosslink reversal; too long = over-fragmentation
- Block age: A 10-year increase in specimen age associates with 39× lower average read depth and 6% lower target coverage. However, 80% of specimens stored 22–32 years still yield successful WES results
- Tissue heterogeneity: Necrotic regions and low tumor cellularity compound fixation problems — poorly preserved areas introduce DNA quality variation that downstream QC cannot fully correct
Applying a Fixed Extraction Protocol Regardless of Sample Variables
Tissue type, section thickness, tumor content, and block age all require protocol adjustments. Labs that apply rigid protocols without QC feedback loops accumulate silent failures that only become visible at the sequencing stage, at which point rework costs (re-extraction, re-library prep, re-sequencing) are highest and sample material may already be depleted.
The downstream impact is significant:
- Preanalytical QC failure rates for clinical FFPE samples intended for comprehensive genomic profiling run 25–40%, primarily due to degraded DNA or insufficient quantity
- In clinical trial cohorts, hybrid-capture CGP test failure rates reach 30–50%
How Cambrian Bioworks Can Help
Manta by Cambrian Bioworks is a clinical FFPE DNA extraction system designed specifically for oncology diagnostic workflows. It holds CE-IVD certification under EU IVDR 2017/746 and ISO 13485 certification for quality management in medical device design and manufacturing — addressing the DNA fragmentation, yield, and reproducibility challenges that define FFPE sample processing.
Key Technical Advantages
- Magnetic bead-based purification recovers fragmented FFPE DNA more effectively than silica column methods
- 28-minute extraction after lysis keeps turnaround tight without sacrificing quality
- Processes 1 to 32 samples per run — no batching pressure for urgent cases
- Pre-filled cartridges reduce manual steps and deliver consistent, reproducible results
- Built-in UVC lamp minimizes cross-contamination risk between runs
- Compact footprint (43.5 cm × 28.2 cm × 26.2 cm) fits inside a standard biosafety hood

Operational Benefits for Clinical Labs
Urgent cancer cases can be processed immediately upon receipt without waiting to fill a batch. Whether handling a single precious biopsy or a full run of 32 samples, the system delivers the same level of automation and reproducibility. Remote monitoring enables real-time run tracking, and the touchscreen interface keeps protocol selection straightforward for routine lab operation.
Proven Track Record in Oncology
Tata Memorial Hospital, ACTREC, and Strand Life Sciences rely on Manta for cancer diagnostics workflows — labs where FFPE extraction quality directly shapes clinical decisions. Cambrian Bioworks designs and manufactures all products in-house in Bangalore, India, and serves labs across India, Europe, the UK, and Singapore.
Frequently Asked Questions
Frequently Asked Questions
What does FFPE tissue stand for?
FFPE stands for Formalin-Fixed Paraffin-Embedded — a preservation method where tissue is fixed in formalin to prevent decay and embedded in paraffin wax for long-term storage. This makes it the global standard for tumor biopsy archiving in clinical pathology.
What are the 4 steps of NGS?
The four core NGS workflow stages are:
- Library preparation: fragmenting and adapting DNA for sequencing
- Cluster generation: amplifying the sequencing template
- Sequencing: reading nucleotide sequences in parallel
- Data analysis: aligning reads and calling variants
Why is NGS better than Sanger sequencing for tumor mutation analysis?
NGS simultaneously analyses hundreds to thousands of genes in a single run at much higher sensitivity. This enables detection of low-frequency mutations in heterogeneous tumor samples — something Sanger sequencing cannot match, given its single-gene, lower-sensitivity approach.
What is the NGS test for tumors?
NGS-based tumor testing, often called Comprehensive Genomic Profiling (CGP), analyses tumor DNA from FFPE biopsies to identify somatic mutations, copy number variants, and structural variants. It also detects biomarkers like TMB, MSI, and HRD that guide targeted therapy selection.
What does a positive NGS test mean for a cancer patient?
A positive NGS result identifies one or more actionable genomic alterations in the tumor, such as a mutation in BRCA1/2, EGFR, or ALK. These findings predict response to specific targeted therapies or immunotherapies, helping oncologists select treatments with the strongest evidence of benefit for that patient.


