Introduction
For labs processing dozens to hundreds of samples daily, DNA extraction from whole blood is one of the most demanding steps in any genomic workflow. Hemoglobin inhibition, variable white blood cell counts, anticoagulant interference, and inconsistent yields compound quickly at scale — and clinical reporting deadlines don't wait.
Managing these variables manually across high sample volumes strains staff bandwidth and introduces the very inconsistencies that compromise downstream assay performance.
That's where the manual vs. automated extraction decision becomes consequential. The choice directly affects cost per test, sample integrity, and whether a lab can realistically scale. With the molecular diagnostics market projected to grow from USD 18.85 billion in 2025 to USD 25.59 billion by 2033 — and blood, serum, and plasma accounting for the largest sample segment — labs that get this decision right are better positioned to grow capacity without proportionally growing costs or errors.
TLDR
- Manual kits offer flexibility and low upfront cost but become labor-intensive and error-prone past 50 samples/day — with limited ROI at scale
- Automation cuts hands-on time by up to 50% and tightens yield consistency: CVs drop below 1%, compared to >15% with manual methods
- Whole blood requires validated protocols to address hemoglobin inhibition, anticoagulant interference, and viscous lysates
- Choice hinges on daily volume, downstream stringency (PCR vs. NGS), and CE-IVD validation requirements
- Automated platforms with CE-IVD certification reduce regulatory burden — a decisive advantage for clinical and diagnostic labs
Automated vs Manual DNA Extraction from Whole Blood: Quick Comparison
| Feature | Manual Spin-Column/Magnetic Kits | Automated Magnetic Bead Platforms |
|---|---|---|
| Typical Platforms | Qiagen QIAamp, Promega ReliaPrep | Thermo KingFisher Flex, Roche MagNA Pure 96, Cambrian Beluga (compact liquid handler) |
| Samples Per Run | 1–24 (centrifuge dependent) | 1–96 samples per run |
| Run Time | 20–40 minutes | 30–90 minutes depending on batch size |
| Hands-On Time | ~6.9 minutes per sample | ~3.3 minutes per sample |
| Typical Yield (200 µL blood) | 3–12 µg | 4–15 µg |
| A260/A280 Purity | 1.7–1.9 | 1.79–1.85 |
| Upfront Investment | Minimal (no instrument) | ₹8,00,000–₹42,00,000+ (approx. $10,000–$50,000 USD) |
| Clinical Validation | IVD kits available | CE-IVD platforms available |

Manual kits excel in flexibility and low barrier to entry, requiring only centrifuges and pipettes. Automated platforms win on reproducibility, walk-away convenience, and scalability. Compact liquid handlers like the Cambrian Beluga (listed above) represent a growing middle ground, giving mid-sized labs automated throughput without the footprint or cost of large-scale dedicated instruments.
What is Manual Kit-Based DNA Extraction from Whole Blood?
Manual kit extraction uses validated reagent kits—typically silica spin-column or magnetic bead formats—where technicians manually perform lysis, binding, washing, and elution. Samples are processed individually or in small batches using multichannel pipettes and 96-well plates.
Why Whole Blood Is Technically Demanding
Whole blood presents specific challenges that complicate manual extraction:
- Hemoglobin inhibition: Hemoglobin and heme are potent PCR inhibitors that reduce amplification efficiency and quench fluorescence-based dyes
- Variable WBC counts: Donor-to-donor white blood cell variability affects yield unpredictably
- Anticoagulant interference: K2EDTA is preferred for genomic applications; heparin inhibits PCR and citrate reduces yields significantly
- Viscosity issues: Lysed blood is viscous and prone to liquid handling errors during manual pipetting
Dominant Manual Kit Chemistries
The two main formats differ in how they capture and wash DNA:
| Kit Format | Mechanism | Scalability |
|---|---|---|
| Silica spin-column (e.g., Qiagen QIAamp) | Chaotropic salt binding to silica membrane; centrifugation for wash and elution | High purity; difficult to scale beyond 24 samples without additional centrifuge capacity |
| Magnetic bead (e.g., Promega ReliaPrep) | Superparamagnetic particles bind DNA via binding buffer; magnetic separation during washing | Scales to 96-well format; compatible with automation when needed |
Use Cases of Manual Kit-Based Extraction
Manual kits fit best in:
- Lower-volume labs (<50 samples/day)
- Research settings with diverse sample types requiring protocol flexibility
- Labs where upfront capital for automation isn't justified
- Rapid turnaround scenarios requiring single-sample extraction
Common downstream applications include targeted PCR panels, SNP genotyping, pharmacogenomics testing, and infectious disease diagnostics. For reference, the Qiagen QIAamp DNA Blood Mini Kit yields 3–12 µg from 200 µL whole blood with A260/A280 ratios of 1.7–1.9—sufficient quality for all of the applications above.
What is Automated DNA Extraction from Whole Blood?
Automated DNA extraction uses robotic liquid handlers or dedicated nucleic acid extraction instruments to perform lysis, binding, washing, and elution with minimal human intervention. Systems process 24–96 samples per run in true walk-away workflows, with some platforms supporting 384 samples per shift.
Magnetic Particle Processing (MPP)
Most automated blood extraction platforms use MPP, where the instrument moves magnetic bead-bound DNA between reagent wells rather than pipetting liquids. The beads are made of superparamagnetic iron oxide particles that exhibit magnetic behaviour only in external magnetic fields, preventing clumping when the field is removed. DNA adsorption to silica bead surfaces occurs via the hydrophobic effect in the presence of chaotropic salts — an entropically driven process that enables efficient capture and release.
Levels of Automation
Semi-automated systems (e.g., Roche MagNA Pure 24) feature compact benchtop instruments with manual plate loading but automated lysis/wash/elute steps. Run times range from 30–60 minutes with less than 5 minutes setup time.
Fully automated systems (e.g., Qiagen QIAsymphony SP) integrate robotic plate handling, reagent dispensing, and barcode tracking. They suit high-volume labs processing hundreds of samples daily.
Critical Advantages for Whole Blood
Automation addresses whole blood-specific challenges that manual processing struggles with:
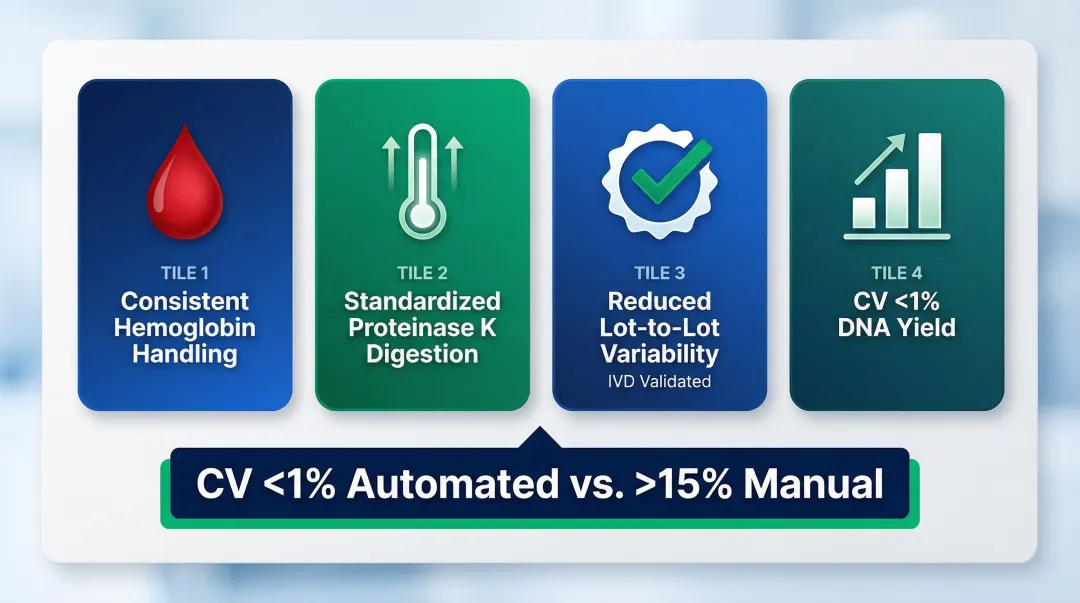
- Handles hemoglobin-rich samples with consistent mixing protocols across all wells
- Delivers standardized proteinase K digestion via on-deck heating modules
- Reduces lot-to-lot variability through validated IVD-grade protocols
- Achieves CV <1% for DNA yield across replicates, versus >15% for manual kits

Cambrian Bioworks' Beluga puts these advantages within reach for labs with limited space. The compact extractor fits inside a biosafety hood, processes 1–32 samples per run without batching pressure, and supports remote monitoring — practical features for busy clinical environments.
Use Cases of Automated DNA Extraction from Whole Blood
These capabilities translate into measurable value across several lab settings:
- Clinical diagnostic labs processing high daily volumes (oncology, prenatal, infectious disease panels)
- Biobanking facilities with large-scale population genomics projects
- Sequencing centres running whole genome or whole exome studies from blood
- Labs under turnaround time pressure for clinical reporting
To put the time savings in perspective: one clinical lab transitioning from manual QIAamp to automated QIAsymphony for Epstein-Barr virus DNA load measurement eliminated 240 pipetting actions and 60 vortex mixing actions per 30-sample batch. That freed up 78 minutes of operator time per run and meaningfully reduced repetitive motion injury risk.
Automated vs Manual Kits: How to Choose the Right Approach for Your Lab
Key Decision Factors
Daily Sample Volume: Automation ROI typically kicks in above 50–100 samples/day. A forensic lab cost-benefit analysis found that automated extraction saved an estimated 107 minutes for 80 samples compared to manual setup.
Downstream Application Stringency: Whole genome sequencing and long-read sequencing demand higher yield and fragment integrity than PCR panels. Automated HMW workflows consistently produce DNA fragments >100 kb for longer read lengths, whereas some manual kits yield fragmented DNA <20 kb — a critical gap for long-read applications.
Staff Availability: Manual extraction requires 6.9 minutes of hands-on time per sample versus 3.3 minutes for automated platforms—compounding significantly at high volumes.
Regulatory Context: IVD-certified workflows are mandatory for in vitro diagnostic reporting in many jurisdictions. CE-IVD marked platforms like Cambrian Bioworks' Manta offer clinical-grade validation for both manual and semi-automated workflows.
The Cost Calculus
The trade-off is straightforward: manual kits cost less upfront but carry higher per-sample labour costs. Automation requires capital investment, but shifts ongoing costs toward consumables — where per-unit pricing is predictable and scalable.
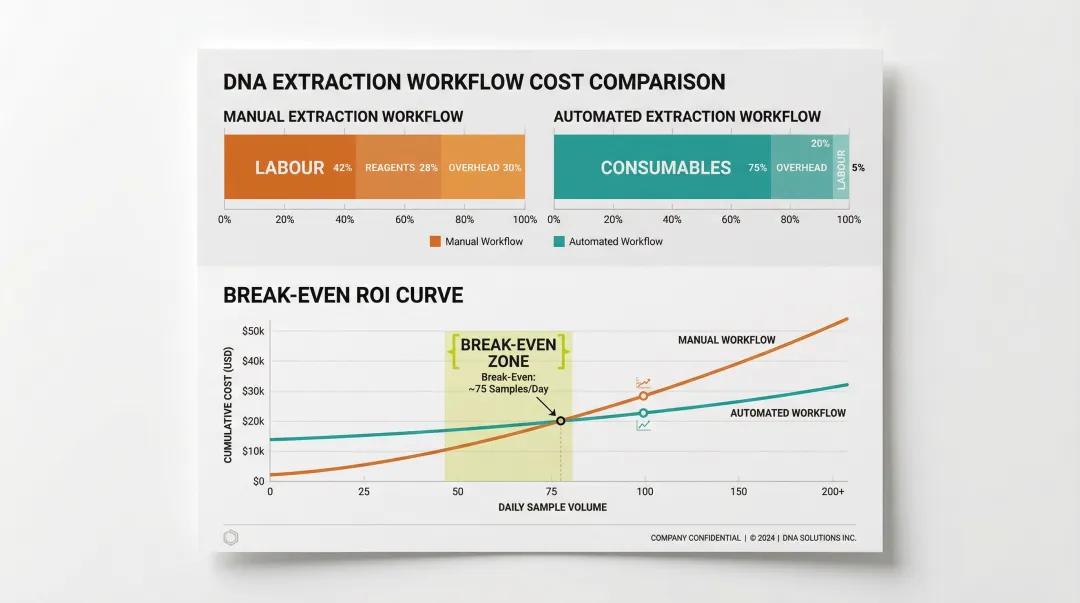
Cost breakdown by approach:
- Manual kits: Labour accounts for ~42% of per-sample cost at volume
- Automated workflows: Labour cost drops sharply; consumables become the primary variable
- Break-even point: Most labs reach automation ROI above 50–100 samples/day
Quality at Scale
At high throughput, manual extraction introduces inter-operator and inter-batch variability that affects downstream assay reproducibility. Automation eliminates this by standardising every extraction step — the coefficient of variation in DNA yield drops to <1% on automated platforms, compared to >15% with manual methods.
Situational Recommendations
Choose manual kits if:
- You process <50 samples/day
- You work with highly variable sample types
- You need protocol flexibility without capital investment
Choose automation if:
- You run high daily volumes
- You require traceable IVD-grade documentation
- You operate with limited staff
- You need to eliminate extraction as a quality bottleneck
What High-Throughput Blood DNA Extraction Looks Like in Practice
A clinical genomics lab processing 80+ whole blood samples daily faced a bottleneck: manual spin-column extraction was causing technician fatigue, and inconsistent yields were affecting downstream NGS library preparation success rates.
After transitioning to an automated magnetic bead-based platform, the lab eliminated 240 pipetting actions and 60 vortex mixing actions per 30-sample batch. Operator time dropped by 78 minutes per run. The coefficient of variation for DNA yield fell from >15% to <1%, improving sequencing library quality and reducing downstream failures.
The trigger for switching wasn't just throughput—it was downstream quality. Inconsistent manual extraction created variability in library prep success rates that only became visible during sequencing data analysis. In most labs, those downstream failures are where the real cost accumulates—not in technician hours logged, but in repeat runs, failed libraries, and delayed reports.
If yield consistency or throughput is creating bottlenecks in your lab, contact Cambrian Bioworks to discuss whether Beluga (automated liquid handler) or Manta (CE-IVD validated extraction kit) fits your workflow.
Conclusion
Manual kits suit flexible, lower-volume labs where capital efficiency matters. For labs where throughput, reproducibility, and clinical-grade traceability are non-negotiable, automated platforms are the clear infrastructure choice. Compact, affordable automated extractors — such as those from Cambrian Bioworks — have made automation accessible beyond large reference labs.
For whole blood specifically, the stakes are high. Hemoglobin inhibition, yield variability, and operator error in manual workflows cascade directly into unreliable diagnostic results. The extraction method you choose affects more than lab efficiency — it shapes the reliability of every result that follows.
Key decision factors to revisit:
- Sample volume and daily throughput — manual works at low scale; automation pays off above ~50 samples/day
- Reproducibility requirements — clinical and regulated settings demand automated consistency
- Budget and footprint — compact automated systems now close the gap for mid-sized labs
Frequently Asked Questions
Is it possible to extract DNA from blood?
Yes, DNA is extracted from white blood cells (leukocytes) present in whole blood. Mammalian red blood cells lack nuclei and do not contribute DNA. Both manual kits and automated platforms reliably isolate genomic DNA from as little as 200 µL of whole blood.
What is a high-throughput DNA extraction method suitable for PCR?
Magnetic bead-based extraction in 96-well plate format—run manually with a multichannel pipette or on an automated liquid handler—is the standard high-throughput approach. It delivers consistent yields, removes inhibitors effectively, and scales well for PCR-ready DNA.
What is a good A260/A280 ratio for DNA?
A ratio of 1.8–2.0 indicates high-purity DNA with minimal protein contamination. Ratios below 1.8 suggest protein or phenol carryover; ratios above 2.0 may indicate RNA contamination—both affecting downstream PCR and sequencing performance.
How much blood is typically needed for DNA extraction?
Most validated protocols require 200 µL to 1 mL of whole blood collected in K2EDTA tubes to yield 3–15 µg of genomic DNA, depending on the donor's white blood cell count and the kit chemistry.
How does hemoglobin affect DNA extraction from whole blood?
Hemoglobin and heme are potent PCR inhibitors that co-purify with DNA when wash steps are insufficient. Whole blood protocols require stringent lysis and multi-step washing—automated systems with consistent mixing cycles reliably produce cleaner eluates than manual processing.
What is the difference between spin column and magnetic bead DNA extraction from blood?
Spin columns use centrifugation to bind DNA to a silica membrane and suit single-sample or small batch processing. Magnetic bead methods use a magnetic field to capture and wash DNA without centrifugation—making them inherently more scalable and compatible with automated liquid handlers for high-throughput applications.


